Dear Editor,
Poikiloderma with neutropenia-Clericuzio type (PNC) is a rare autosomal recessive disorder characterised by widespread post-inflammatory poikiloderma, pronounced neutropenia and frequent sinopulmonary infections.1 It results from mutations in the USB1 gene, with pathogenic variants like c.531delA, c.496delA, c.179delC and c.326_327del reported to date.2-4 Its exact prevalence remains unknown, with a few cases reported in the literature. Herein, we report a PNC case in an adult female patient with a novel mutation in the USB1 gene.
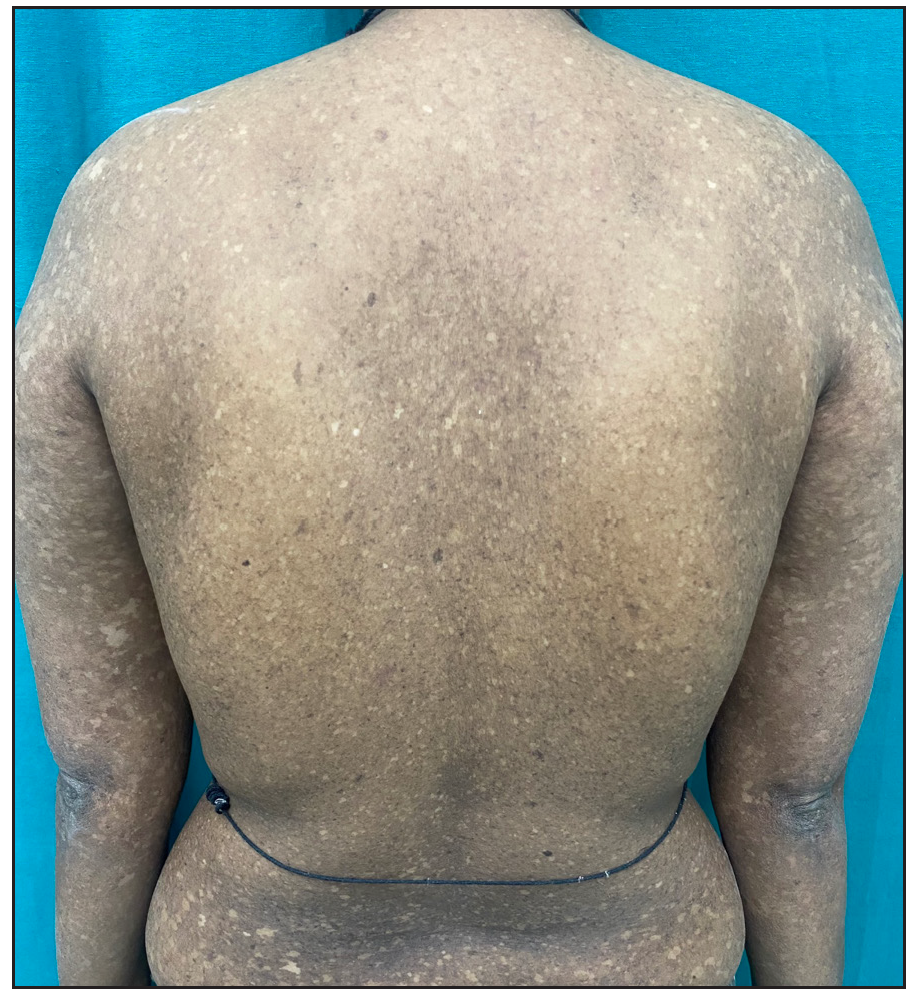
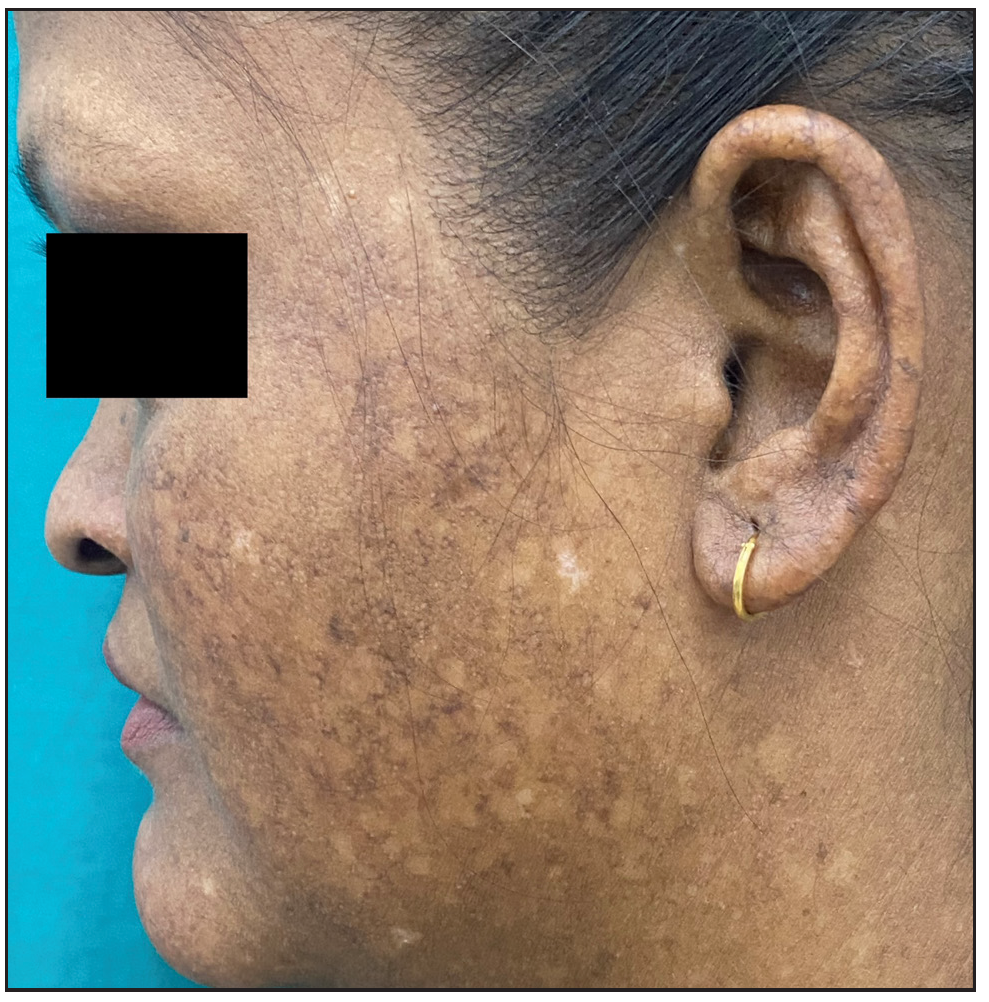
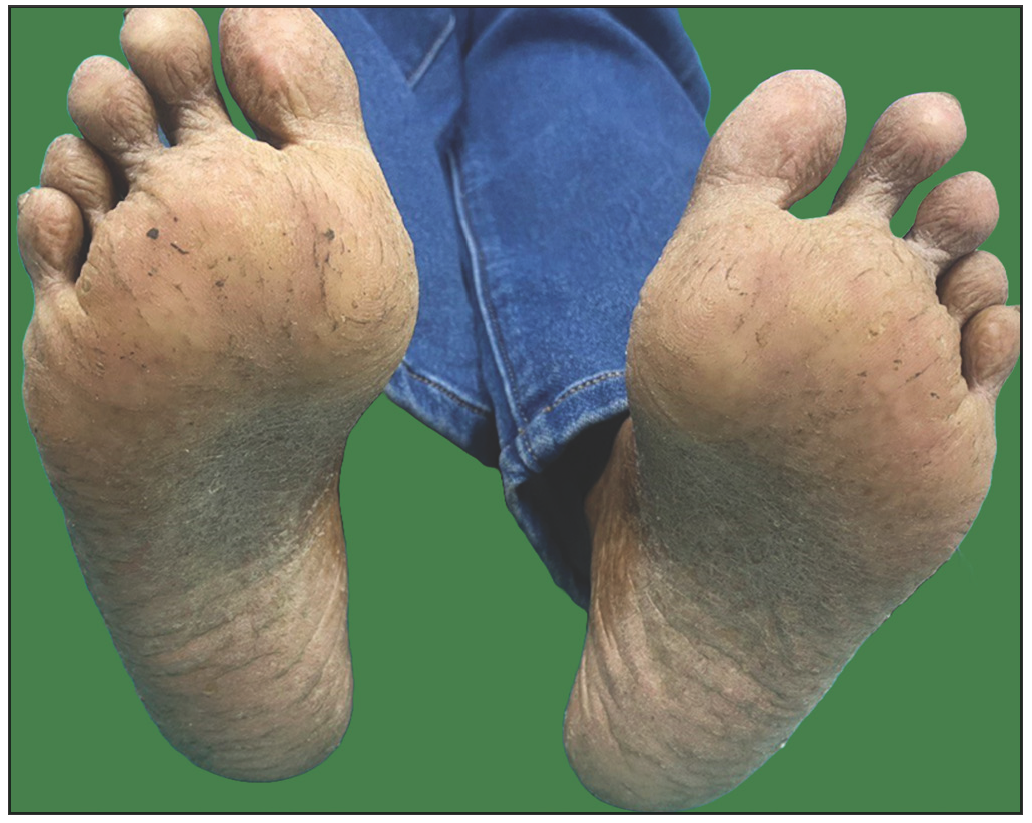
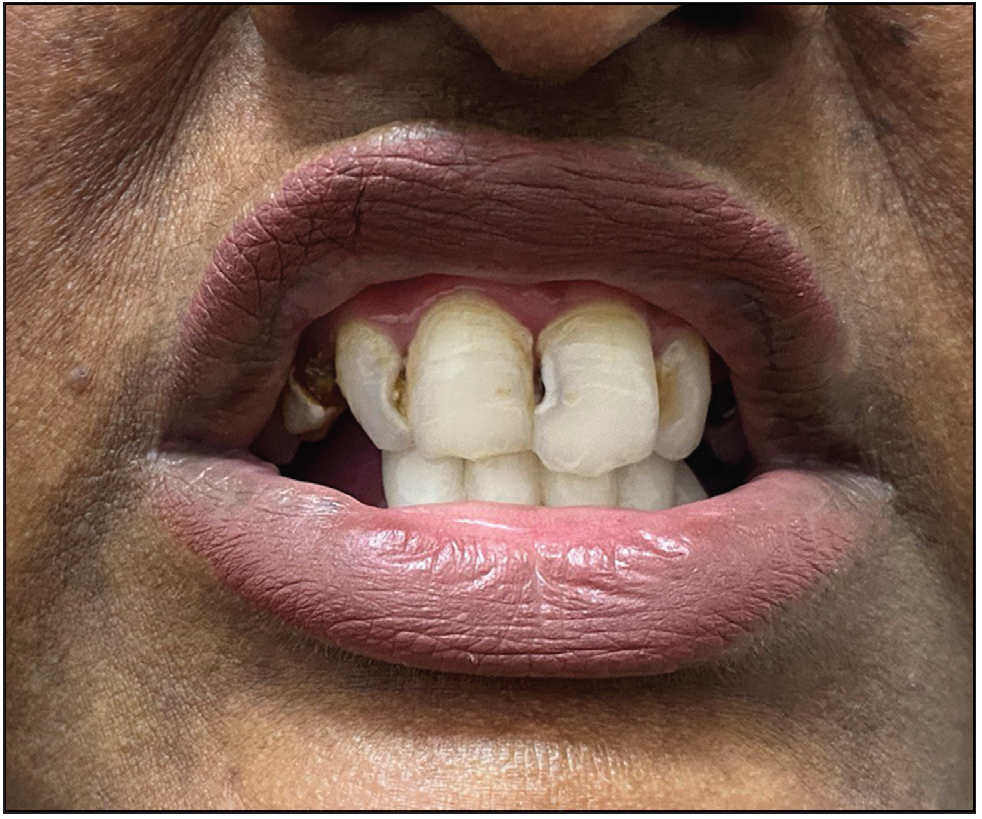
A 26-year-old woman, born to non-consanguineous parents, presented with hyperpigmented and hypopigmented spots all over the body for seven months of age. She had a history of recurrent sinopulmonary infections associated with fever, malaise and lethargy. However, she denied recurrent cutaneous infections such as recurrent pyoderma, abscess, cellulitis, otitis media, bone involvement or non-healing ulcers. No similar complaints were reported in the family. Physical examination showed a prominent forehead, saddle nose and normal weight and height. Cutaneous examination revealed multiple ill-defined, raindrop-like, hypopigmented, atrophic macules on a background of reticular hyperpigmentation on the trunk [Figure 1a], upper and lower extremities and face [Figure 1b]. There was also a patchy loss of eyebrow hair on the lateral half. Plantar keratoderma [Figure 1c] with normal palms and preserved dermatoglyphics were observed. Toenail dystrophy with mild subungual hyperkeratosis was noticed, while fingernails were normal. Oral examination revealed peg-shaped teeth, loss of multiple molars and premolars, and crowding of central incisors and canines [Figure 1d]. Other mucosa and body hair were normal. Laboratory evaluation showed leukopenia (total leukocyte count: 1500/mm3) with neutropenia (380 cells/mm3). Serum ferritin and lactate dehydrogenase levels were elevated. Bone marrow studies showed no significant abnormality. Other systemic evaluations, including chest X-ray were normal. Whole exome sequencing (WES) identified a homozygous 9-base pair deletion in exon7 of the USB1 gene (c.741_749del variant). Based on the above findings, a diagnosis of PNC was made. This variant was classified as a variant of uncertain significance and had not been previously reported in 1000 Genomes, TopMed, ClinVar, gnomAD, human gene mutation database or online mendelian inheritance in man databases. However, considering the size of the mutation (9 base pair inframe deletion), homozygosity and genotype-phenotype correlation, it was assumed to likely be a novel pathogenic mutation. The patient and her family were counselled for Sanger sequencing and segregation analysis; however, they declined due to financial constraints. She was briefed about the long-term prognosis and advised stringent photo-protection as well as topical keratolytics for plantar hyperkeratosis. Annual complete blood count and dermatological and pulmonary surveillance were advised along with a bi-annual dental examination. Patient was counselled for encouraging at-risk relatives for screening to ensure early diagnosis and prevention of complications.
Export to PPT
Export to PPT
Export to PPT
Export to PPT
PNC must be differentiated from other poikilodermatous disorders, such as Rothmund Thompson syndrome (RTS), Kindler syndrome and dyskeratosis congenita (DKC).5 PNC is characterised by widespread poikiloderma, recurrent infections and cyclic neutropenia due to the mutations in the USB1 gene, which encodes an exonuclease critical for U6 snRNA maturation and spliceosome assembly.3,6 This mutation leads to defective snRNA processing, disrupts haematopoiesis and increases the risk of myelodysplastic syndrome (MDS).3 DKC presents with poikiloderma, oral leukoplakia and dysplastic nails but lacks neutropenia, distinguishing it from PNC. Studies show that PNC can lead to global bone marrow dysfunction affecting all three haematopoietic lineages, increasing the risk of progression to MDS and in some instances, acute myelogenous leukaemia.3 Similarly, individuals with DKC are at an increased risk for bone marrow failure and skin cancers.7 RTS, another close differential, is characterised by poikiloderma, sparse hair, dental and skeletal abnormalities, and an elevated malignancy risk, especially for osteosarcoma and cutaneous malignancies.5 A notable characteristic of RTS is the specific pattern of poikiloderma that begins on the face and extends to the extremities while sparing the trunk. In PNC, it initiates peripherally before progressing to the face and trunk. Another distinguishing trait of RTS is the presence of radial ray abnormalities along with a normal neutrophil count. Managing PNC necessitates a multidisciplinary approach, involving dermatologists, haematologists, geneticists and other specialists for comprehensive care, close surveillance and timely interventions. This case emphasises the importance of revisiting diagnoses in congenital photosensitivity disorders given their considerable phenotypic overlap, thereby highlighting the need of genetic testing for accurate diagnosis. Our report is limited by the absence of in silico analysis and inability to perform Sanger sequencing and segregation analysis in family members for confirming the mutation. However, the identification of a novel variant in the USB1 gene is significant and underscores the ongoing need of genetic research in rare disorders.
Comments (0)