In nature, bacterial communities display both individual and collective behavior. Collective behaviors are the result of coordination of activity between many individual cells. Signal exchange is one mechanism by which bacteria coordinate collective behaviors. Signal exchange between bacteria is often referred to as quorum sensing (QS), as the concentration of signaling molecules, called autoinducers, is a proxy to measure cell population density [1, 2]. Synthase enzymes produce signals, and receptor proteins detect these signals and respond by regulating gene expression within the cell [3]. A large change in gene expression, and thereby the activity state of a population of bacteria, is enacted when the concentration of a molecular signal exceeds a threshold. In this sense, cells utilizing QS can transition between activeand inactive states, with active states being associated with dense populations of cells at high signal concentrations. Due to positive feedback in many QS systems, specifically the increased expression of the proteins that produce and detect molecular signals, the transition from QS inactive to active state is sharp and is often approximated as a two-state system [2, 4, 5].
QS is typically studied as a process utilized by one species of bacteria to regulate its own activity state. However, in many natural contexts, many bacterial species coexist within diverse communities of cells [6–8]. Within such communities, multiple species of bacteria utilize homologous signaling pathways, one example being multiple species producing and sensing acyl-homoserine lactones (AHLs). Receptors for these signals can be promiscuous, interacting with multiple variants of a signal molecule and resulting in crosstalk [9]. Here, we define crosstalk as an interaction between signaling molecules and receptors from different QS pathways, in which the combination of signals impacts the activation or repression of gene expression. This definition aligns with prior literature [10, 11]. Crosstalk has also been used in the context of distinct two-component systems influencing the same cellular response [12]. The main focus of this study is on what has been called interspecies crosstalk [10],the result of multiple species of bacteria utilizing homologous signaling pathways. Such crosstalk has been known for many years and has been both reported in experiments and analyzed theoretically [9–11]. Crosstalk can either activate or repress changes in gene expression to a variable degree, depending on the identity of both the signal molecule and receptor protein [13]. Recently, a model was proposed and validated that incorporated these variable crosstalk interactions [14]. Crosstalk was examined within communities of Bacillus subtilis that exchanged up to 5 different variations of the ComX peptide signal, and a basic neural network model accurately predicted the activity state in such networks [15].
One unexplored context of signaling within diverse bacterial communities is the exchange of multiple, orthogonal signal variants. Orthogonal signals are sets of signals that are detected by distinct receptor proteins. A specific pair of signal molecules is considered orthogonal if the first signal molecule does not influence the cellular response to the second signal molecule, and vice versa. Each signaling molecular has a separate and independent signal transduction pathway. Examples of orthogonal sets of bacterial signals would be AHLs, autoinducing peptides (AIP), and autoinducer-2 (AI-2). A set of signaling molecules is a group of one or more chemically similar molecular signals that interact with a common receptor protein. For example the receptor LuxR from Vibrio fischeri detects and responds to the 3-oxo C6 HSL produced by other Vibrio fischeri cells, but also detects and responds to other AHLs signals produced by other bacterial species, such as C8-HSL, 3-oxo-C12 HSL or C4-HSL [9, 10, 16]. LuxR will not detect or respond to AIP or autoinducer-2 [17], making AHLs one distinct set of signals. However, some species of bacteria are well known to have multiple receptor proteins that detect distinct sets of signals. The best studied example is Vibrio harveyi that uses LuxN, CqsS, and LuxPQ receptors to detect the HAI-1, CAI-1, and AI-2 AIs receptively [18, 19]. This is one of several such species that utilize multiple QS systems [10, 17, 20–22]. Species that detect multiple AIs often regulate a similar set of genes in response to any of these individual signals. In other words, cells with receptors for two orthogonal sets of signaling molecules detected by separate receptors will transition to the active state in response to high concentrations of either signal [23, 24]. Here, we assume that activation through one signaling pathway results in activation of all signaling pathways within a species. These cells act as ‘or’ logic gates, with any one set of signals leading to activation of two separate signaling pathways and overproduction of both associated signals. It is unclear how this coupling of orthogonal signaling networks impacts the regulation and stability of QS-regulated activity states within bacterial communities.
In this work we build upon a neural network model, inspired by the Hopfield model of neurons [14, 25, 26], to investigate the emergent properties of signaling networks within diverse bacterial communities. Within the model, a population of a bacterial species is represented by a node, and edges between the nodes represent signal crosstalk. Communities of cells using sets of orthogonal signals are represented as multilayer networks, with bacterial species able to detect and respond to orthogonal sets of signals being common nodes that connect these layers. We use this modeling framework to predict the activity state of the community and explore to what extent these activity states are stable to perturbations. Further, we identify the features that determine the resilience of activity states in QS communities, and analyze the potential for perturbations to spread from layer to layer. This analysis will help develop strategies to design and manipulate the activity states of multispecies microbial communities regulated by the exchange of complex mixtures of signaling molecules.
The model, shown in figure 1(A), illustrates a network with three species of bacteria that exchange a common set of QS signal variants. Each species is represented by a node of the network. The size of the nodes represents the population size of each species. Each bacterial species produces one chemical signal, but potentially detects multiple signals being produced within the community. This framework is inspired by the ability of QS signaling pathways to be affected by non-cognate but chemically similar signals produced by other species or strains of bacteria, such as AHL crosstalk in Gram-negative bacteria and AIP crosstalk in Gram-positive bacteria [13, 27]. The pairwise-weights between nodes represent the QS crosstalk interactions between the species, with the sign of the weight indicating whether the specific signal molecule promotes or inhibits QS activation in a particular species. Thus, whether a species transitions to being QS active depends on the weighted concentrations of all signals in the network. The change in signal concentration is modeled using an ODE,
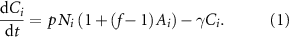
Figure 1. A neural network model of signal exchange in diverse microbial communities. (A) In the model, each species is represented by a node labeled a, b, or c. The size of the node represents the population of each species. Each species produces a unique signal molecule and arrows represent the influence of signal exchange on node activity (red arrows show negative interactions and green arrows positive interactions). The numbers on the arrows represent the sampled weight values Wij, which quantify the strength and sign of crosstalk between nodes. Each node is either in the quorum sensing off state (represented by 0) or the quorum sensing on state (represented by 1). (B) A simulation of signaling dynamics. Upon accumulating signal concentrations above the threshold, dashed horizontal line, nodes a and b activate. Active nodes have a larger rate of signal production, leading to a jump in signal concentration. The chart indicates the initial (i) and final (f) QS activity state of each node. (C) Simulations examined the role of crosstalk, comparing networks with non-zero weights connecting nodes to a network with crosstalk weights of zero. For networks with 2–20 nodes, there was an increase in the fraction of active nodes for networks with crosstalk. Error bars show the standard error from 2000 simulations. The p-value calculated using Friedman ANOVA test was <10−100 when comparing No crosstalk with Crosstalk.
Download figure:
Standard image High-resolution imageHere p is the production rate of signal, γ is the degradation rate of the signal, 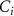 is the concentration of signal i,
is the concentration of signal i, 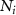 is the population of bacterial species i,
is the population of bacterial species i, 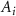 represents the activity state of node i with 0 being inactive and 1 being active. Upon activating, the production rate of signal increases from p to fp, where f is the fold change in the expression of the signal producing enzyme.
represents the activity state of node i with 0 being inactive and 1 being active. Upon activating, the production rate of signal increases from p to fp, where f is the fold change in the expression of the signal producing enzyme.
Change in the activity of any node is based on whether or not the weighted sum of signals exceeded a threshold concentration, following:
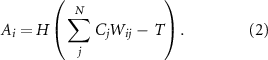
Here, Wij is the weight representing how signal produced by node j influences the activity of bacteria in node i, T is the threshold concentration of signal, and H is the Heaviside function. Activity takes values of 0 or 1. The weight matrix, W, is such that Wij represents the influence of signal from the jth node on QS activity state of cells in the ith node. The weighted sum includes all signals produced by all N nodes within the system, including the signal Ci produced by node i. Weights between nodes, 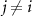 , were sampled from a normal distribution with mean 0 and standard deviation 0.4 (figure S1(A)). Wii was set to 1.0, representing a positive influence of a node’s cognate signal on its own activity.
, were sampled from a normal distribution with mean 0 and standard deviation 0.4 (figure S1(A)). Wii was set to 1.0, representing a positive influence of a node’s cognate signal on its own activity.
Figure 1(A) shows a three species network with arrows representing the crosstalk weight matrix. The model focuses on how changes in the population size and signal weights impact the activity state of the network, as these two terms define the community composition, i.e. the bacteria species within the community and their relative abundances. Changes in population sizes would have the same consequence as a proportional change in signal production rate. There are similar connections between the fold change, signal degradation rate, and population size, but we chose not to lump multiple variables together to highlight the impact of changes in the community composition. Other variables might also change due to evolution of the genes in the signaling pathway, but that was not the focus of this study.
The change in signal concentration and activity states of the above network following these equations are shown in figure 1(B). At time t = 0, all nodes were in the off state and signal concentrations were 0. Over time, signals were produced and the concentrations increased. Nodes activated if the weighted sum of input signals to that node exceeded the threshold signal concentration (equation (2)). In figure 1(B), nodes ‘a’ and ‘b’ activated near time = 20, and node ‘c’ remained in the inactive state. Upon activation, because signal production increased by the fold change factor f, the signal concentration increased at a faster rate. Note that the dashed line shows the threshold concentration of total weighted signal required for activation, while the curves show the concentration of individual signals. Therefore, the jump in signal production sometimes occurred when a signal was slightly above or below the dashed threshold line. Eventually signal concentrations reached steady-state values, as first-order signal degradation eventually balanced signal production.
2.1. Crosstalk increases activity within the networkIn figure 1(C), we investigated how signal crosstalk impacted QS activation for networks with 2–20 nodes. Activity was compared for networks with and without crosstalk between species. Networks with crosstalk, as shown in figure 1(A), have interaction weights between nodes sampled from a normal distribution with mean 0 and standard deviation 0.4 (figure S1(A)). For the no-crosstalk case, all weights between nodes are zero, indicating that the activity state of each node is solely dependent on the signal produced by that node and the population size of that node. For both contexts, the self-weight, Wii, was set to 1.0 for each node. Each condition was simulated 2000 times. For each iteration, the population sizes and interaction weights were randomly rechosen. Population sizes were chosen from a distribution described in the methods, such that the average size of a node is 100 cells and the total population of all nodes is 100 times the number of nodes (figure S1(B)). Each simulation was run for 200 time steps, which as shown in figure S2 was sufficient for networks to reach steady-state signal concentrations. The fraction of active nodes after the final time step was recorded and plotted against the network size in figure 1(C). The average fraction of active nodes in the crosstalk and no-crosstalk cases saturated at 60% and 40% respectively. This indicated that crosstalk promotes higher average network activity compared to networks where nodes do not interact with each other. For the no-crosstalk case, the decrease in the average fraction of active nodes as the number of nodes increased was related to a small shift in the distribution of node sizes for networks with more nodes, see figures S1(C) and (D).
The model makes a few strong assumptions that warrant some discussion. First, population sizes did not change over time steps as cell replication was not included in the simulation. No growth approximated the context of a population with relatively constant structure over long periods of time. Figure S3 explores networks that include cell growth. Changes in QS activity are made at high cell density as signal concentrations begin to approach threshold concentrations for a QS response. Therefore, model results including the transient period of cell growth while the network is accumulating but not yet responding to released signals did not noticeably change the activity states of the networks or the response to perturbations. For more sophisticated and realistic models with different growth parameters for each species, the order of node activation and phenotypic memory within QS may influence final activity states of the community [28], although these cases are likely atypical. The model presented here represents the simplest case without growth. Second, the average weight assigned to a crosstalk interaction was 0, with an equal probability to negative and positive weights. Prior work has shown both negative and positive crosstalk weights between bacteria [13, 27, 29]. To assess the impact of biased crosstalk interactions, we varied the mean of the sampled weight distribution. Networks with a positive mean had significantly higher average activity states, while those with negative mean showed suppression. This supports the idea that even slight biases in signal crosstalk can reshape network-wide activity (figure S4).
2.2. Stability of the activity state to perturbationsTo examine the stability of these networks, changes of the network activity state to perturbations in signal concentration or population size were analyzed. As shown in figure 2(B), initially signal concentrations were set to zero with all nodes in the inactive state. Over time, signal concentration and node activity states changed according to the equations (1) and (2). After 200 time points, the activity state of the network was determined. Then, a perturbation, defined as a change in the signal concentration or population size of a node, was applied. The perturbation caused an instantaneous change in one signal concentration or population size. The system then responded to the perturbation, by producing signal and potentially updating node activity states, for an additional 200 time points. To quantify changes in the network activity state 200 time steps after the perturbation, the hamming distance (HD) was calculated:
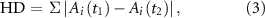
Figure 2. Stability of bacterial signaling networks to perturbations. (A) Initially all nodes begin in the inactive state and then over time the network approaches a stable activity state. The network is then perturbed and given time to reach a stable activity state. Calculation of the normalized hamming distance quantifies how many nodes changed activity state after the perturbation. (B) Example of applying a perturbation to a 3-node network. After time t = 200, an 80% decrease in the concentration of signal a resulted in the activation of node c. (C), (D) Quantifying the robustness of a 10-node network to perturbations in signal or population size. Targeted perturbations are applied to the node with the largest population size. Random perturbations are applied to a random node. Error bars show standard error from 2000 simulations. p < 10−100 when comparing random vs. targeted data in (C) and (D), calculated using Friedman ANOVA test.
Download figure:
Standard image High-resolution imagewhere Ai is the activity state of node i, t1 is the time just before the perturbation and t2 is the time at which the network state is re-evaluated after the perturbation, after 200 additional time steps. The number of nodes that flipped their steady-state activity state post-perturbation is defined as the HD. A HD of 0 indicates that the network’s activity state before and a long time after the perturbation is the same. This tells us that the network was robust to the applied perturbation. On the other hand, a HD > 0 indicates one or more nodes changed activity state. The normalized HD (NHD) is the HD divided by the total number of nodes in the network. Figure 2(B) depicts the response of an example 3-node network to a perturbation. After 200 timepoints, the concentration of the three signals reached a steady-state, with two nodes being active and one node being inactive. At time point t = 201, there was an instantaneous 80% decrease in the concentration of signal ‘a’. As the network responded to the perturbation, the concentration of signal ‘a’ returned to the unperturbed level while the previously inactive node ‘c’ activated and remained in the active state. Node ‘b’ remained in the active state. Hence, the perturbation led to a change in the activity of one node.
To generalize the behavior of such networks, figure 2(C) explores the response of many networks to a range of perturbations. Here, each network had 10 nodes. For each realization, the crosstalk weights and node population sizes were re-sampled as described in Methods. Perturbations of signal concentration were between −100% to +100%. Perturbations of population sizes were −90% to +100%. It should be noted that population sizes did not change following a perturbation to a node population size, in other words changes in the size of a node were permanent. Perturbations to random and targeted nodes were compared, denoted by green and purple curves respectively. Targeted perturbations were only applied to the largest population size in the network and random perturbations were applied to a random node, which includes the possibility of the random node being the largest population size. A sample of 2000 networks were analyzed for each perturbation size. Targeted perturbations exhibited a higher mean NHD than a random node perturbation (figure 2(C)). As an example, an NHD of 0.1 means that for the 10-node network simulated, 1 node changed activity state. Figure 2(C) also revealed that perturbing the population of a node resulted on average in a higher NHD than perturbing the signal concentration.
2.3. Extension of the model to multilayer signaling networksAs previously mentioned, several species of bacteria have been identified which utilize multiple orthogonal QS signals. As a point of definition, orthogonal signals are signaling molecules that do not interact with the same receptor protein. For example, some Gram-negative bacteria exchange both homoserine lactones and AI-2, but AI-2 does not directly bind to or interact with the HSL receptor and vice versa. Often a similar set of genes is differentially expressed in response to high concentrations of either signal. To capture the potential for some species to exchange two different sets of orthogonal signals, a multilayer network model was developed.
This multilayer network model is two separate single layer models with one or more shared nodes, as depicted in figure 3(A). Any node in both layers of the network is referred to as a common node. Common nodes detect and produce signals in both layers of the network, and the activity state of a common node is calculated as an OR function. In other words, if the signals in either layer of the network result in activation, the node was activated in both layers. The node will produce signals in both layers at the activated rate. Nodes that are only found in one layer cannot directly communicate with nodes in the other layer, but the common nodes can relay information between layers of the network through changes in activity state.
Figure 3. A model of multilayer bacterial signaling networks. (A) For networks that utilize two orthogonal sets of signals, a multilayer model was developed. Each layer of the model consisted of a group of species that exchanged a common set of signals. If one species detected both sets of signals, that node existed in both layers of the network (node ‘a’ in the multilayer network connected with the blue line). Activation in one layer translates that activity to the second layer. (B), (C) The response of two-layer networks to perturbations of one of the signal concentrations. These networks have 10-nodes per layer with 0, 1, or 2 common nodes. (B) reports the change in the activity state of the perturbed layer, while (C) reports the change in the activity state of the unperturbed layer. Networks with 0, 1, or 2 common nodes were compared using Friedman ANOVA test. P-values for data in (B) and (C) are <10−8 and <10−88 respectively. Error bars show in (B) and (C) show standard error for 2000 replicate simulations.
Download figure:
Standard image High-resolution imageThis ‘OR’ logic is motivated by biological precedent. In Vibrio harveyi, three distinct quorum-sensing systems (AI-1, AI-2, and CAI-1) converge on the master regulator LuxR, such that activation through any pathway can drive the same regulon [18, 24]. Vibrio fischeri has a similar gene network architecture, as measurements of AI-2 and two different AHLs are integrated to determine QS activity [30]. In Pseudomonas aeruginosa, the las, rhl, and PQS pathways co-regulate overlapping sets of genes and act in a hierarchical fashion, effectively coupling distinct signals to common outputs [21]. Recent synthetic systems further demonstrate that distinct quorum-sensing modules can be engineered to jointly control a single circuit, highlighting the feasibility of OR-like integration [16]. The assumption that QS activation is coupled between pathways utilizing orthogonal sets of signals is a strong assumption, and it allows the coupling within the multilayer networks. Alternative logic structures, such as additive or fractional-threshold rules, could be explored in future work to capture systems where signal pathways regulate partially overlapping or distinct regulons.
The response of such two-layer networks to perturbations was explored. To determine if the number of common nodes impacted stability of the multilayer network to perturbations, networks with 0, 1, or 2 common nodes were compared. These two-layer networks had 10 nodes in each layer. The size of each node was assigned as described in Methods, and for a common node the population size in the first layer was copied to the second layer.
As in figure 2, the network evolved starting from zero concentrations of all signals at t = 0. After 200 time steps, a perturbation of a signal concentration from one of the nodes was applied. The perturbed signal is produced by one of the nodes in the network, and the layer containing this node is labeled as the perturbed layer. The layer that does not contain the node associated with the perturbed signal is labeled as the unperturbed layer. Common nodes, existing in both layers of the network, were not perturbed.
The network then evolves for 200 additional time steps, and NHD was used to quantify changes in the activity state of the network after the perturbation. As shown in figure 3(B), stronger perturbation caused larger changes in the activity state. Figure 3(C) analyzes activity changes in the unperturbed layer, revealing that the perturbation had a much stronger impact on activity in the perturbed layer as compared to the unperturbed layer, although in rare cases the activity state of nodes changed in the unperturbed layer. The impact of the perturbation on the activity of the unperturbed layer depended on the number of common nodes. As expected, with no common nodes the perturbation did not change the state of the unperturbed layer. Networks with 2 common nodes had a NHD that was more than 2-fold larger than networks with 1 common code. The response of two-layer networks to perturbations in population size is shown in figure S5, revealing similar trends. The number of nodes that flip their activity states following a signal perturbation in two-layer networks with 0, 1, or 2 common nodes is shown in figure S6 and table S1. The plots reveal that following a 90% signal perturbation, >85% of the time no nodes changed state and occasionally up to 6 nodes on the perturbed layer and 2 nodes on the unperturbed layer changed activity state.
2.4. Common nodes correlate the activity states of orthogonal signaling networksTo further quantify the degree to which perturbations of one layer influence activity in the other layer, mutual information (MI) was calculated. MI between the perturbed and unperturbed layer NHD’s is given by equation (4).
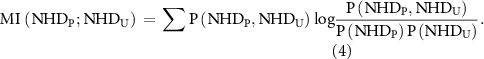
Here, NHDP and NHDU are the observed NHD’s representing the NHD in the perturbed and unperturbed layers, respectively. 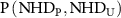 gives the joint frequency of observing NHD in perturbed layer and NHD in unperturbed layer, and
gives the joint frequency of observing NHD in perturbed layer and NHD in unperturbed layer, and 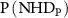 and
and 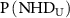 are their corresponding marginal distributions. MI is the measure of independence between the joint NHD of the two layers and the product of marginal NHD distribution of each network. A value of 0 indicates that the probability distribution of the NHDs of both networks are independent, while a value >0 indicates that the probability distributions of NHDs of the two networks are correlated. The higher the MI, the more correlated the networks.
are their corresponding marginal distributions. MI is the measure of independence between the joint NHD of the two layers and the product of marginal NHD distribution of each network. A value of 0 indicates that the probability distribution of the NHDs of both networks are independent, while a value >0 indicates that the probability distributions of NHDs of the two networks are correlated. The higher the MI, the more correlated the networks.
When either increasing the common nodes or perturbation strength, MI increased (figures 4(A) and (B)). This indicates that changes in the activity of the two networks were more correlated with more common nodes. This analysis was done for both signal perturbations (figure 4(A)) and population size perturbation (figure 4(B)). Similar trends in the mean NHD with respect to perturbation size were produced for both types of perturbations, although MI increased with stronger perturbations (figures 4(A) and (B)). We also performed such perturbation analysis on the common nodes as shown in figures S7 and S8 and found them to be equally important for the transmission of perturbations between the layers.
Figure 4. Factors that determine the sensitivity of networks to perturbations. (A) The mutual information quantifies the correlations in the activity state of two 10-node networks before and after perturbations of one signal concentration. Signals associated with a random node were perturbed, excluding common nodes. Mutual information for two-layer networks with 0, 1, or 2 common nodes was compared. (B) Mutual information calculated for perturbations of population size. (C), (E) Response of a two-layer network to perturbations in the signal associated with nodes of different size. Nodes were ranked by relative size in the network (C) or grouped by absolute size (E). (D), (F) Response of a two-layer network to perturbations in the signal associated with nodes of different interaction strengths, quantified as the sum of the absolute value of all outgoing weights at each node. Nodes were ranked by relative (D) or absolute (F) sum of outgoing weights. The average NHD of both layers is plotted against the relative and absolute values of these quantities in figures (C)–(F). Error bars show the standard error of NHD with each condition being repeated in 10 000 networks for (C) and (D) and 1000 networks for (E) and (F). For each iteration, networks were reassigned population sizes and weights. In all plots, the blue line shows networks with 0 common nodes, red is 1 common node, and orange is 2 common nodes. P-values comparing networks with 0, 1, or 2 common nodes in panels (C)–(F) were calculated using the Friedman ANOVA test, giving <10−30, <10−30, 0.0233, and 0.3629, respectively.
Download figure:
Standard image High-resolution image 2.5. Large nodes are most influential in setting network activity statesNext, we quantified the influence of each node on the overall community activity state. This analysis sought to reveal which node characteristics were most influential in setting the network activity state. Nodes have two key properties, the population size and the set of interactions weights between nodes. The response of a network to perturbation of a single node was measured by calculating the NHD.
Common nodes were not perturbed in this analysis. The NHD was compared when nodes of different population sizes and weighted interactions were perturbed. The population size of a node is straightforward. The magnitude of the interactions at a given node was assessed by taking the sum of the absolute value of all outgoing weights. The relative rank of these two values was assigned as compared to the other 20 nodes in the same network. The average NHD is plotted against the relative and absolute values of these quantities in figures 4(C)–(F).
Figure 4(C) shows that the perturbation of larger nodes is more likely to result in a change of network state, particularly if the largest population size is perturbed. Figure 4(E) shows that nodes with 300–500 cells have the largest NHD. This is in agreement with figure 4(C), as the largest population size in the network typically has >200 cells, figure S9. Surprisingly, the perturbation of a node with more than 600+ cells has a reduced likelihood of a change in network state as compared to a node with 500–599 cells. Because a 10-node network cannot have two nodes with more than 500 cells, this implies that perturbation of smaller largest nodes results in a larger chance of a change in the network state. Largest nodes with population sizes >600 produce a larger fraction of the signal within the networks, as compared to largest nodes with smaller population sizes. Therefore, nodes with population sizes >600 likely reactivate following the perturbation and the signal from this node strongly influences the network activity state. The signal contribution from smaller nodes is too ‘weak’ to influence the final network activity state following perturbation. For networks with ‘smaller’ largest nodes, the size difference between the first and second largest nodes is likely smaller. Network activity states are more equally determined by the signal from several nodes of similar size. Transient activity state changes following perturbation are less likely to reset to the pre-perturbation activity state, hence a higher NHD.
Conversely, the absolute sum of interaction weights at a given node did not predict the NHD following perturbation. The NHD was not dependent on the magnitude of interaction weights with other nodes in the network. These simulations were repeated using a perturbation of population size instead of signal concentration, figure S10, showing similar trends. Weaker perturbations were also applied, as shown in figure S11, revealing that the strength of the perturbation could influence trends in which population sizes were most impactful to network resilience.
Understanding how cell signaling impacts the regulation of cellular activity states is a critical problem in biology. Although significant prior work has examined cell signaling from the viewpoint of systems with one or two bacterial species and one or two signals [31–33], signaling within more complex systems, with multiple bacterial species and multiple signals, remains less understood. Here, the ability of signal produced by one bacterial species to influence the response of a different bacterial species to its own, cognate signal is referred to as crosstalk. Biological crosstalk has also been used in another context, to describe interactions between distinct receptor proteins to modulate a common response regulator [12]. Wu et al also discuss both signal crosstalk and promoter crosstalk in the context of bacterial QS [11]. A definition from Sanders et al describes crosstalk as ‘any mixing between two signaling pathways A and B that have their own signal inputs and outputs, but wherein signal A also modulates to some extent the output of receptor B, and vice versa’[16]. This definition applies well to both the external signal exchange between bacteria and the internal mixing of information stemming from two separate sensors. Here the term crosstalk is applied to the context of a bacterial species with a receptor that responds to a mixture of two or more distinct, external molecular signals. In many natural contexts, including signaling within diverse bacterial communities, the exchange of dozens of chemical signals is expected [3, 34, 35]. To help understand signaling within such contexts, a neural network model of signaling in bacterial communities was previously reported [14, 15]. We build from this prior work to explore the response of such networks to perturbations and to include signaling networks that produce more than one set of signals. Here, networks can produce orthogonal sets of signaling molecules that do not directly interact. Such situations are expected to be common in natural contexts, as sets of orthogonal signals such as AHL, AIP, AI-2, and other signals have been reported [36]. Within each set of signals, variable crosstalk between pairs of signals and receptors was included. Because some species of bacteria utilize more than one set of signals [3, 24], there is the potential to transfer activity between signaling networks. Such common nodes lead to coupling between the activity states regulated by signals that do not directly interact. The NHD quantifies this coupling by counting the number of nodes in the network that change states in response to a perturbation. Though we found it convenient to carry out our analysis of node importance via NHD measurements, a linear response matrix is also a useful tool for this task [37].
This model has produced several key insights into signaling within diverse bacterial communities. First, the presence of crosstalk in QS networks leads to more activity within the network. This increase in activity was shown for networks with an average weight of zero. Even with many negative weights within the network, 20% more nodes reach the active state. An explanation for this observation is that activity states are strongly influenced by the largest population size, and the largest population size tends to be in the active state. The active large node will have positive crosstalk with about 50% of the other nodes in the network, leading to the activation of nodes that are above or near the activation threshold. This observation may be relevant to signaling within natural ecosystems of cells, as crosstalk in general, even if sometimes negative, plays a role in increasing QS activation. Network activity should also mirror the pattern of crosstalk weights associated with the largest population size of the network.
Another insight is that the largest nodes have the most impact on network activity state. As shown in figure 2, targeted perturbation of the largest node in the network had a two-fold increase in the HD. The importance of the node with the largest population was reinforced by the figure 4(C), revealing that the perturbation of the largest node resulted in a much larger increase in the probability of a change in activity state as compared to when other nodes were perturbed. This result is not surprising, given that a node’s influence on the activity within the network is dependent on signal concentration, and signal concentration is proportional to population size. Larger nodes produce more signal. Figure 4(E) highlights that networks that are least robust to perturbations are ones with ‘smaller’ largest population size. If the largest node is more than 600 cells (representing about 30% of the total population), networks are more resilient to perturbations. This is because the large node recovers from the perturbation and re-establishes the original activity state. However, when the largest node is smaller in size, that implies the network contains a few large nodes of similar size. Such networks have a greater probability of transitioning to a new, steady-state activity state following a perturbation.
Overall, multispecies signaling networks are also largely robust to perturbations of either signal or population. The average normalized HD s in network activity state are only up to 20% for the most extreme perturbation that target the largest node, which indicates only 2 nodes changing activity state within a 10-node network. For weaker perturbations, the normalized average HD of the network activity state is close to 0.02 or 0.01, which indicates on average 0 or 1 nodes within a 10-node network change activity state following perturbation. Node removal was also examined in figure S12, showing that removal of the largest node in the network had a NHD above 0.2. Here we considered perturbations of only one network component. Potentially perturbations of many signals or populations at once would lead to larger changes in the network activity state.
An intriguing finding is that nodes that utilize two orthogonal signals, common nodes, have the potential to transmit perturbations between two orthogonal signaling networks. As shown in figure 3(C), perturbations to signal in one layer of a two-layer network have a small chance of leading to changes in state of the second layer that was not directly perturbed. The normalized HD in this case is small, but this result highlights the important although weak coupling between orthogonal signaling networks. This result suggests that manipulation of one set of signals in a bacterial community could have unanticipated consequences in orthogonal signaling states. There is also the potential to control the activity state of a species of interest indirectly, through perturbations of a signal that is not recognized by the target species. Given the structure of the networks analyzed, any change in the activity state of a node in the unperturbed layer would require a change in the activity state of a common node. For −90% signal perturbation causing an NHD > 0 in the unperturbed layer, 20% of cases showed that nodes other than common nodes also changed activity state (figure S13). The perturbation of common nodes directly, as shown in figures S7 and S8, showed a response similar to a perturbation applied to a random node.
A key practical reason for understanding the robustness of activity states within heterogeneous QS networks is to control QS-regulated phenotypes within heterogeneous bacterial communities. Naga et al reviewed various strategies of antimicrobial therapy via QS inhibition, amongst which include disrupting the signaling systems [38]. Jiang et al explained QS receptor inactivation as a technique in fighting bacterial diseases [39]. This effectively is represented by a population size perturbation in our model, as removing cells permanently stops the production of AI from those cells. Signal degradation by enzymes is another method, which corresponds to a signal perturbation in our model, although potentially a permanent decrease [40, 41]. Such strategies may be effective, although prior work has largely ignored the system-level consequences of manipulating a specific component of the communication network. In contexts in which regulation of QS-regulated phenotypes is a goal, such as P. aeruginosa in the lungs of cystic fibrosis patients [42], the target species lives in a heterogeneous community with the potential for signal crosstalk between species. Pathogens, including P. aeruginosa, also utilize multiple, orthogonal signaling pathways [43]. Such species that utilize multiple signals not only can respond to multiple sets of signals, but also play an important role in relaying activity information between groups of species that do not directly communicate. In these contexts, analyzing community-level signaling should be beneficial to the regulation of QS-regulated genes within heterogeneous populations of bacteria. Such analysis will reveal the potential for indirect regulation of signaling states and strategies for effectively controlling the activity state of a target species.
4.1. Distributions of population size and weightsThe population size of each node is sampled from a modified negative exponential distribution, as shown in figure S1(B). This modification, done to better match real microbial communities, which have many species with small populations and a few species with large populations [44, 45]. This distribution is realized by choosing boundaries at random between 1 and the largest numbers of cells and using the length of the line segment to populate each node. For example, in a network having 5 nodes with an average population size of 100 cells the total cell population is 500. 4 random values are chosen from the uniform distribution between 1 to 499. The random integers are arranged from smallest to largest. The difference between the lowest integer and 0, consecutive values of these integers, or the total population size and the largest integer provides the values for the cell-population of each node. This creates a population distribution following a modified negative exponential. In the case of two-layer networks with common nodes, the total population of both the networks is distributed among N—C nodes, where N is the total number of nodes in both layers of the network and C is the number of common nodes.
The weights of a network are captured in a matrix of size N × N, where N is the total number of nodes of a network. For networks with cross-talk, the elements of the network are sampled from a normal distribution with a mean of 0 and variance of 0.4, as shown in figure S1(A). This ensures an equal probability of sampling positive and negative weights. The diagonal elements are set equal to 1, which indicates species self talk is positive and has a strong influence on activation. For networks with no cross-talk (figure 1(C)) all non-diagonal elements of the cross-talk matrix are set to 0, with the diagonal elements remaining at 1.
4.2. Threshold signal concentration for QSNodes change state based on a Heaviside function that compares the weighted sum of signals to the threshold signal concentration. Thus, in order for node i to activate, the following must be satisfied:
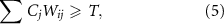
where again, Ci is the signal concentration associated with node i, Wij is the crosstalk weight from node j affecting node i, and T is the threshold signal concentrati
Comments (0)