The emergence of antibiotic resistance, which often occurs under changing levels of antibiotics is a major concern for human health [1, 2]. In an important class of antibiotics, called bacteriostatic antibiotics [3], the drug does not induce death directly, but renders some essential process in the cell metabolism less efficient or inactive [4–9] resulting in a reduced cell growth. For these antibiotics, it is essential to properly model cell metabolism and cell growth in order to better understand the action of antibiotics [10–13]. According to [14], the distinction between bacteriostatic and bactericidal antibiotics that target ribosomes depends on the value of their dissociation rate from the ribosomes. Antibiotics with slow dissociation rates are more likely to induce cell death and be classified instead as bactericidal because of the depletion of essential proteins they cause.
In the field of bacterial growth, the study of growth laws [15–17] represents a major step forward in our understanding of cell growth. These growth laws result from conservation of ribosome capacity and flux balance at steady-state. Recently, a new way to understand them has been put forward, that relies on a description of the cell metabolism as an ensemble of autocatalytic cycles, such as the cycle of ribosome production and that of RNA polymerase production [18]. By definition, an autocatalytic cycle implies a set of reactions and a set of species, so that all species of the set can be produced in increasing number by running the autocatalytic cycle [19]. The method, which we apply in this paper, has wide applications for addressing various questions related to cell growth [20]. For instance, it has been recently used to formulate predictions about the interplay between cellular growth rate and mRNA abundances [21].
While some predictions about the action of RNA-polymerase targeting antibiotics have been derived in [18], the full consequences for the inhibition of growth by a general antibiotics have not. In particular, this work does not discuss the second growth law that describes the inhibition of translation by antibiotics, nor the possibility that there may be two different regimes for the action of antibiotics, namely the so-called reversible and irreversible binding regimes of antibiotics. This distinction is quite important in practice because for reversible binding, faster growth in the absence of the drug leads to an increased susceptibility, while the opposite is true for irreversible binding [13]. Further, a regime of antibiotics concentration exists where two values of growth rate are possible (growth rate heterogeneity or bistability [22]) for the same range of physical parameters. A recent study of a wide class of antibiotics reported that this growth rate heterogeneity is variable depending on the target molecules [23], but at the moment, it is not clear how to model or predict this growth rate heterogeneity.
To summarize, we believe that the inhibition of bacterial growth by antibiotics has not been considered from a sufficiently general point of view, which motivates the present study. By building on references [18] and [13, 24], we develop a general framework for the inhibition of bacterial growth by bacteriostatic antibiotics in which the cell metabolism is modeled as coupled autocatalytic cycles. Our work applies to all antibiotics for which the target molecule is an intermediate in an autocatalytic cycle of the cell metabolism that couples to cell growth, leaving aside other types of antibiotics that may for instance kill the cell without affecting its growth. Given the central role played by [13, 24] in our work, we start by a quick summary of the main findings of this paper. In the next section, we present our model, so that the new elements which we have introduced appear clearly. Then, we explore some consequences, concerning growth laws, and we test our model with experimental data on the dependence of the growth rate as function of the concentration of antibiotics for a wide range of different antibiotics. Then, we introduce a new proxy of cell risk induced by the antibiotics and we present some extensions of our model for more complex situations involving the combined action of multiple antibiotics [25–27] or the indirect effect due to the accumulation of a product with inhibition properties.
Here, we recapitulate the main findings of a classic model of inhibition of bacterial growth by antibiotics [13], which is applicable to antibiotics that target ribosomes. In this model, the cell is viewed as a compartment in which the antibiotic present outside the cell can enter and bind to ribosomes. The perturbation of translation produced by the antibiotics is described by growth laws, which quantify the interdependence of the cell growth rate λ with the intracellular ribosome concentration r. The first law states that the ribosome concentration should increase linearly with the growth rate according to [15–17]:
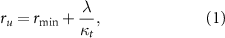
where κt, ru and 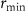 are respectively the translation capacity, the concentration of unperturbed ribosomes and a minimal ribosome concentration.
are respectively the translation capacity, the concentration of unperturbed ribosomes and a minimal ribosome concentration.
The second growth law states that, in the presence of an antibiotic inhibiting translation, the ribosome production is up-regulated, which also leads to another linear relation
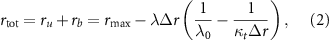
where where rb is the concentration of ribosomes bound to the antibiotics and 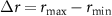 is the dynamic range of the ribosome concentration. In other words, the second growth law describes the increased production of ribosomes that follows translation inhibition. As a result, the total ribosome concentration becomes negatively correlated with the bacterial growth rate in the presence of these inhibitors.
is the dynamic range of the ribosome concentration. In other words, the second growth law describes the increased production of ribosomes that follows translation inhibition. As a result, the total ribosome concentration becomes negatively correlated with the bacterial growth rate in the presence of these inhibitors.
Now the antibiotics enter the cell and bind to the ribosomes, with the rate 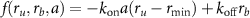 where
where 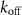 and
and 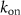 are first and second order rate constants and a is the antibiotic concentration inside the cell. Antibiotics can only bind to ribosomes if their concentration is above the minimum threshold
are first and second order rate constants and a is the antibiotic concentration inside the cell. Antibiotics can only bind to ribosomes if their concentration is above the minimum threshold 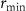 according to this formula. The flux of antibiotic concentration into the cell is
according to this formula. The flux of antibiotic concentration into the cell is 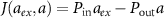 , where aext is the antibiotic concentration outside the cell. Antibiotics enter the cell with rate
, where aext is the antibiotic concentration outside the cell. Antibiotics enter the cell with rate 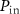 and exit with rate
and exit with rate 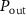 , which could occur due to diffusion by passive transport or through pores by active transport [28, 29].
, which could occur due to diffusion by passive transport or through pores by active transport [28, 29].
Together, these assumptions lead to the following dynamical equations [13]:
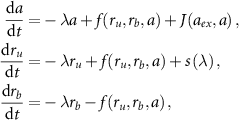
where 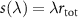 represents the ribosome synthesis rate.
represents the ribosome synthesis rate.
In the absence of inhibitors, the pre-exposure or basal growth rate is λ0, which corresponds to the normal behavior of the cell. The steady-state solution of this model is given by the following cubic equation [13]:
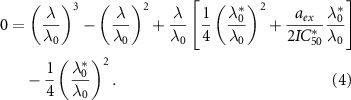
The reversibility of the binding of the antibiotic is characterized by the parameter 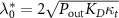 , KD is the dissociation constant
, KD is the dissociation constant 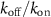 and
and 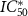 is a typical concentration such that
is a typical concentration such that 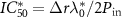 . Since equation (4) is a cubic equation in the growth rate, there are one or three solutions, and in particular there is a parameter regime in which the dynamical system can be multi-valued.
. Since equation (4) is a cubic equation in the growth rate, there are one or three solutions, and in particular there is a parameter regime in which the dynamical system can be multi-valued.
The model predicts two regimes depending on the value of 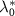 , called the reversible and irreversible limits. The reversible limit
, called the reversible and irreversible limits. The reversible limit 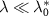 describes a regime of strong outflux of toxic agents and unbinding rate. In that case, the growth rate has a smooth behavior described by:
describes a regime of strong outflux of toxic agents and unbinding rate. In that case, the growth rate has a smooth behavior described by:
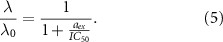
This smooth behavior is due physically to a rapid equilibrium which is reached between intra and extra cellular antibiotic pools.
In contrast, the irreversible limit 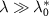 corresponds to negligible outflux and unbinding rate compared to the influx of toxic agents and binding rate. In that case, one obtains a discontinuous function:
corresponds to negligible outflux and unbinding rate compared to the influx of toxic agents and binding rate. In that case, one obtains a discontinuous function:
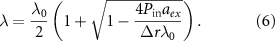
In this regime, the system behaves as a toggle switch behavior, due to the competition between the antibiotic influx and the ribosome production.
By analyzing various types of antibiotics, the authors of [13] found that experimental data for bacteriostatic antibiotics indeed fit into one class or the other. Another major insight of the model, was the prediction of different growth dependent susceptibility for the two classes of antibiotic behaviors. This susceptibility is measured thanks to the half-inhibition concentration IC50, which is defined as the concentration of toxic agent at which the growth rate is half its initial value. This is a measure of the sensitivity of the system to external stress, the higher it is, the more resistant is the system to inhibitors. By substituting 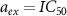 and
and 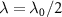 into equation (4), one finds that the half inhibitory concentration IC50 falls onto a universal growth dependent susceptibility curve:
into equation (4), one finds that the half inhibitory concentration IC50 falls onto a universal growth dependent susceptibility curve:
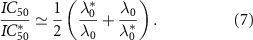
We now introduce our model for cell metabolism as two coupled autocatalytic cycles, in which one cycle describes the production of ribosomes, while the other describes RNA-polymerase production [18]. These two autocatalytic cycles are coupled because ribosomes are necessary to synthesize RNA-polymerase protein subunits and vice-versa for ribosomes: B1 represents the number of active ribosomes; C1 the number of active RNA polymerases; similarly 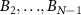 and
and 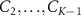 are the abundances of intermediates involved in the assembly of ribosomes and RNA polymerases respectively, BN; CK are the abundances of fully assembled but resting ribosomes/RNA polymerases respectively. Similarly to the classic model we first presented, we suppose that ‘toxic’ inhibiting agents in numbers A can bind to one of the autocatalysts (chosen here to be B1 for simplicity) with a rate
are the abundances of intermediates involved in the assembly of ribosomes and RNA polymerases respectively, BN; CK are the abundances of fully assembled but resting ribosomes/RNA polymerases respectively. Similarly to the classic model we first presented, we suppose that ‘toxic’ inhibiting agents in numbers A can bind to one of the autocatalysts (chosen here to be B1 for simplicity) with a rate 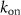 and unbind with a rate
and unbind with a rate 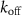 , proportionally to the relative abundance of antibiotics in the cell [13]. We denote
, proportionally to the relative abundance of antibiotics in the cell [13]. We denote 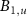 the abundance of unbound ribosomes and
the abundance of unbound ribosomes and 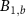 the abundance of bound ribosomes.
the abundance of bound ribosomes.
The signification of the different variables in the model is summarized in the table 1.
Table 1. Variables of the model. Note that we used dimensionless numbers for species within the cell, except for aex which has the unit of a concentration and Ω which has the unit of a volume.
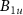 Number of fully formed free active ribosomes
Number of fully formed free active ribosomes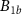 Number of fully formed ribosomes bound to antibioticsANumber of toxic agent molecules within the cellaexConcentration of toxic agent molecules outside the cellΩCell volumeBk for
Number of fully formed ribosomes bound to antibioticsANumber of toxic agent molecules within the cellaexConcentration of toxic agent molecules outside the cellΩCell volumeBk for 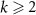 Number of ribosomes precursorsC1Number of fully formed and active RNA-polymerasesCk for
Number of ribosomes precursorsC1Number of fully formed and active RNA-polymerasesCk for 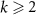 Number of RNA-polymerase precursorsN (resp. K)Number of building steps for ribosomes (resp. RNA-polymerase)
Number of RNA-polymerase precursorsN (resp. K)Number of building steps for ribosomes (resp. RNA-polymerase)In our model, we rely on Leontief’s production function [19, 30] (see appendix section E for details), according to which the rates of reactions involving two complementary resources are set by the limiting quantity among the two using a minimum function. Historically, this law of the minimum has been introduced by Leontief’s in his work in economy [31]: in a network of firms producing one product each by consuming the outputs of other firms (resources), the rate of production will be set by the availability of the scarcer resource. A similar idea was developed later by Liebig in ecology [32]. More recently, it was used for modeling autocatalytic cycles in metabolism [18]. With this method, we get linear equations in regimes where one reactant is scarce. This is similar to assuming that one reactant is in excess in a chemical reaction, and that the kinetics is set by the concentration of the scarcer reactant.
Unlike in the previous model, which was formulated in terms of concentrations, our approach uses abundances or numbers [33] precisely because it is based on the Leontief framework. Due to this difference, our dynamical equations formulated in terms of species numbers do not contain the growth rate explicitly unlike equation (3). Naturally, it is straightforward to show that the two formulations are equivalent, because of the assumption of balanced growth for the cell. In this regime, all species present in the autocatalytic cycles grow at the same rate 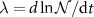 , where
, where 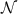 is typically the number of ribosomes or RNA-polymerases... Note that the cell volume is not constant but also grows at the same rate as the abundances of species inside the cell. This is the reason for the use of fractions defined with respect to the total abundances of mature molecules
is typically the number of ribosomes or RNA-polymerases... Note that the cell volume is not constant but also grows at the same rate as the abundances of species inside the cell. This is the reason for the use of fractions defined with respect to the total abundances of mature molecules 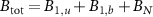 .
.
One can then combine the equations of the model to obtain a linear matrix equation for the sub-populations of ribosomes only, without explicit dependence on antibiotics, and a self consistent equation for the growth rate λ of the whole cycle (see appendix, section A). In the following, we assume the cycle targeted by the toxic agent becomes limiting. The effect of the inhibition of one cycle on the other cycle is only considered in appendix, section C.4. Consequently, we isolate the inhibited cycle and study its growth, because it restricts the growth of the rest of the network.
A key quantity is the fraction of active ribosomes 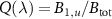 , which takes the form of a polynomial in terms of the cell growth rate λ with factors depending on the rate constants
, which takes the form of a polynomial in terms of the cell growth rate λ with factors depending on the rate constants 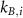 of reaction steps of the autocatalytic cycle :
of reaction steps of the autocatalytic cycle :
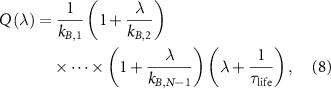
where 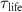 is the life time of mature intermediates BN,
is the life time of mature intermediates BN, 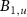 and
and 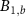 , which corresponds to the time of degradation of these molecules. This life time is assumed to be of the same order for all these species for simplicity and is typically large in comparison with the growth rate, unless the cell is in a regime of reduced growth [21].
, which corresponds to the time of degradation of these molecules. This life time is assumed to be of the same order for all these species for simplicity and is typically large in comparison with the growth rate, unless the cell is in a regime of reduced growth [21].
The expression above simplifies to 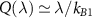 in the limit of ‘fast assembly’
in the limit of ‘fast assembly’ 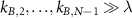 and long ribosome lifetime
and long ribosome lifetime 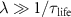 . In this case, the results do not depend on the number of steps N in the first cycle. We also understand from equation (8) that if one step n becomes limiting, the term
. In this case, the results do not depend on the number of steps N in the first cycle. We also understand from equation (8) that if one step n becomes limiting, the term 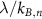 cannot be ignored, which modifies
cannot be ignored, which modifies 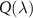 . With the above conditions, we recover the linear increase of the fraction of unbound ribosomes with respect to λ, which is the first growth law:
. With the above conditions, we recover the linear increase of the fraction of unbound ribosomes with respect to λ, which is the first growth law:
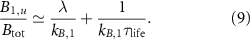
Note that this is the equivalent of equation (1) in the previous model. This law describes the increase of the fraction of unbound ribosomes with the growth rate under changes of nutrient quality in the absence of antibiotics, so when 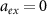 . Here, increasing the nutrient quality can be realized by increasing assembly rates
. Here, increasing the nutrient quality can be realized by increasing assembly rates 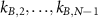 , assuming that they are equal to each other. Indeed, if only one of these rates were increased, the other steps would be limiting and we would not see the effect we are interested in. In the end, we obtain the solid blue curve in figure 1(b), which approaches the origin when λ goes to zero due to the long lifetime assumption.
, assuming that they are equal to each other. Indeed, if only one of these rates were increased, the other steps would be limiting and we would not see the effect we are interested in. In the end, we obtain the solid blue curve in figure 1(b), which approaches the origin when λ goes to zero due to the long lifetime assumption.
Figure 1. (a) Scheme of coupled autocatalytic networks interacting with a toxic agent. The blue box linking two arrows represents a coupling through a min function [31, 32]. (b) The first growth law is the increase of the ribosome fraction with the growth rate (solid curve), the second growth law corresponds to the dashed colored lines obtained by varying the amount of antibiotics. The pre-exposure growth rate λ0 displayed on the right scale.
Download figure:
Standard image High-resolution imageWhen an antibiotic inhibiting translation is present, the ribosome fraction 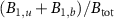 decreases with the growth rate, which is the second growth law [15]. With our formalism, we indeed obtain a negative correlation between these variables, which takes a linear form:
decreases with the growth rate, which is the second growth law [15]. With our formalism, we indeed obtain a negative correlation between these variables, which takes a linear form:
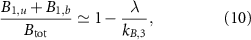
if we assume fast assembly, fast activation, long ribosome lifetime 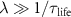 and a single intermediate step (N = 3). This equation is the equivalent of the second growth law described by equation (2). Without specific assumptions on the rates, one obtains the colored curves in figure 1(b), which have been obtained by varying the external concentration of antibiotics aex keeping all other parameters fixed.
and a single intermediate step (N = 3). This equation is the equivalent of the second growth law described by equation (2). Without specific assumptions on the rates, one obtains the colored curves in figure 1(b), which have been obtained by varying the external concentration of antibiotics aex keeping all other parameters fixed.
As the concentration of antibiotics increases, the growth rate always decreases below the basal growth rate λ0. We find that for 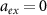 , the decreasing and increasing curves of figure 1(b) cross each other, which is expected because
, the decreasing and increasing curves of figure 1(b) cross each other, which is expected because 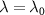 at this point.
at this point.
It is important to appreciate that the first and the second growth laws are derived from our model, while they were introduced as phenomenological constraints in equations (1) and (2) in the model we first presented. Further, in the original work on growth laws [17], linear dependencies with respect to the growth rate were reported. In contrast to this, we see from figure 1, that neither the first nor the second growth law is strictly described by linear relations. A small curvature is indeed visible for the curves that correspond to the second growth law (when the translation in inhibited by an antibiotic) in the experimental data of [17], which is reported in figure 5 of [34], yet the few experimental points make it complicated to assess this effect clearly. It is difficult to appraise whether there is a curvature for the first growth law in this data, because there is not enough data available in the low growth rate region.
We now explore further consequences of our formalism. Let us first consider the case of arbitrary number of intermediates (N), for which we have obtained the self-consistent equation for 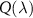 given in equation (34).
given in equation (34).
The reversible limit 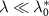 describes a regime of strong outflux of toxic agents and unbinding rate. We find that in this limit (see appendix, section A):
describes a regime of strong outflux of toxic agents and unbinding rate. We find that in this limit (see appendix, section A):
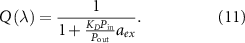
In contrast, the irreversible limit 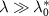 corresponds to negligible outflux and unbinding rate compared to the influx of toxic agents and binding rate. Then, we obtain a different equation setting the growth rate (see appendix, section A):
corresponds to negligible outflux and unbinding rate compared to the influx of toxic agents and binding rate. Then, we obtain a different equation setting the growth rate (see appendix, section A):
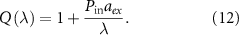
For ribosomes in the regime of intermediate or high growth rates, we can expect a long lifetime, a small resting rate, fast assembly and fast activation [18]. These conditions translate to 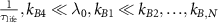 , yielding
, yielding 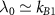 . In this limit, we can simplify our self-consistent equation for the growth rate
. In this limit, we can simplify our self-consistent equation for the growth rate
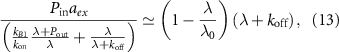
so that we recover the equation derived in [13]. With the additional assumption of fast binding 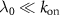 , the possible values of the growth rate are roots of a polynomial, from which it is possible to recover the reversible and irreversible limits of antibiotics binding described previously. Further, we find in this limit
, the possible values of the growth rate are roots of a polynomial, from which it is possible to recover the reversible and irreversible limits of antibiotics binding described previously. Further, we find in this limit 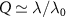 .
.
Interestingly, the self-consistent equation for the growth rate obtained within the autocatalytic framework (see appendix, section A) has two solutions in the irreversible limit with fast assembly, leading to two separate branches of solutions for λ. A first solution remains close to 0, corresponding to a non-growing cell. A second one is larger but exists only until a given concentration of inhibitors is reached, above which the system jumps on the other branch, and the growth rate vanishes as shown in figure 4(a). In experiments, in the irreversible case, the system usually starts from λ0 and the growth rate decreases as the concentration of inhibitors increases, until the discontinuity where the growth rate jumps on the second branch and vanishes. This growth rate heterogeneity happens above a threshold in terms of the antibiotic concentration. Such a phenomenon has been predicted in other theoretical works [13, 35], and it has also been observed experimentally [22, 36].
A very recent experimental study by Kals et al quantified the growth heterogeneity (GRH) systematically for many antibiotics across different concentrations [23]. In this work, the growth rate heterogeneity is evaluated from the standard deviation of the growth rates at a given time point and averaged across multiple time points. The authors reported that this GRH typically increases near the minimum inhibitory concentration and can be large or small depending on the type of antibiotics. In agreement with our predictions, they observed a small GRH for chloramphenicol and a large GRH for ampicillin or rifampicine for instance. One should note however, that in our model, the distinction between reversible or irreversible, which controls the GRH, is not purely a matter of the target molecule of the antibiotic. According to the definition of 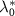 , the size of the efflux rate
, the size of the efflux rate 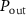 matters also. This means that a molecule like tetracycline, which is expected to be reversible and to have a small GRH [23], can nevertheless display a coexistence between growing and non-growing states in some conditions as observed experimentally, when the duration of the drug exposure is short and in the presence of additional regulation mechanism that controls the efflux rate [37].
matters also. This means that a molecule like tetracycline, which is expected to be reversible and to have a small GRH [23], can nevertheless display a coexistence between growing and non-growing states in some conditions as observed experimentally, when the duration of the drug exposure is short and in the presence of additional regulation mechanism that controls the efflux rate [37].
We have tested our model on a number of antibiotics, for which experimental data can be found in the literature [9, 13]: Chloramphenicol inhibits ribosome production by binding to ribosomes, preventing them from transcribing new proteins; Rifampicin targets RNA-polymerase by binding to RNA-polymerase [38, 39]; Kanamycin, Streptomycin, Chloramphenicol and Erythromycin target the ribosomal autocatalytic cycle [4, 6, 8, 40]; and finally Triclosan targets the synthesis of fatty acids [41–43], thus affecting the building of bacterial membranes [18].
In figure 2, we show the normalized growth rate 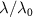 as function of the concentration of antibiotics only for Chloramphenicol and Kanamycin, the plots for the other antibiotics are shown in appendix, section B. The details of the fitting procedure, as well as the constraints and values imposed for the different parameters are explained and justified in appendix section B.2, where a table sums up these results.
as function of the concentration of antibiotics only for Chloramphenicol and Kanamycin, the plots for the other antibiotics are shown in appendix, section B. The details of the fitting procedure, as well as the constraints and values imposed for the different parameters are explained and justified in appendix section B.2, where a table sums up these results.
Figure 2. Comparison with experiments for two drugs affecting bacterial growth, namely (a) Chloramphenicol (data from [9, 13]) and (b) Kanamycin (data from [13]). The solid line shows the growth rate as a function of the fraction of inhibitors, while the dotted line shows a measure of the risk faced by the cell defined in the text. The data were fitted by constraining the parameters as explained in appendix. Different experiments for the same antibiotic correspond to different growth medium.
Download figure:
Standard image High-resolution imageIn [18], the effects of Triclosan and Rifampicin were explained by adding Hill functions heuristically to describe saturation effects in the cycle. In contrast here, we provide an explicit expression for the dependence of the growth rate on the fraction of antibiotics without such an assumption. The fact that we are able to describe a large panel of antibiotics suggests that these antibiotics can indeed be depicted as inhibitors affecting essential cellular autocatalytic cycles despite their different mechanisms. Note that we recover different concavities in figure 2, which correspond to the two distinct regimes of cellular response to the antibiotics previously identified for ribosome-targeting antibiotics [13]: the reversible limit where the outflux of antibiotics compensates the influx of the latter, and the irreversible limit where antibiotics bind quickly to autocatalysts, resulting in an accumulation of bound, inhibited individuals.
Antibiotics have a rather limited number of targets such as ribosomes [3, 4, 6, 10, 13] or RNA-polymerase [7] for instance. Regardless of the mechanism of action or precise targets, the effect of antibiotics on growth show similarities [12], which suggests that a general measure of the risk induced by the toxic agent might exist. In particular, bacteriocidal antibiotics do not appear to be fundamentally different from bacteriostatic ones, both reduce cell growth, but if the inhibition is too strong, processes that are necessary for survival cannot be satisfied and cell death can occur [10, 44]. In fact, it has been demonstrated that for ribosome targeting antibiotics, the cidality depends on the rate of dissociation of antibiotics (and thus on the amount of bound antibiotics in the cell) [
Comments (0)