Targeted radionuclide therapy (TRT) has enabled the specific targeting of cancer cells by exploiting the charged particles emitted during the radioactive decay near the cancer target. These charged particles induce DNA damage leading to cancer cell death. However, the nature of the particle influences not only the penetration depth but also the complexity of the resulting cellular damage. While β-emitters like Lutetium-177 have reached the clinic for the treatment of metastatic prostate and neuroendocrine cancers, they may be less effective at treating micrometastases and disseminated tumour cells because a substantial proportion of their disintegration energy escapes and is deposited outside the tumour volume (Champion et al 2008, Hindié et al 2016). Therefore, there is a renewed interest in α-emitters and Auger electron (AE) emitting radionuclides, which have a lower tissue range but a higher linear energy transfer (LET) which enable them to deliver high absorbed doses in small lesions without cross-dose toxicity (Bernhadrt et al 2001, Falzone et al 2015). AEs, originating from an inner electron shell vacancy, are very low-energy electrons with a penetration depth of a few nanometers. The potential of AE-emitting radionuclides for TRT has been well reviewed in literature (Bolcaen et al 2023, Aboagye et al 2023). Due to the very short range of AEs and conversion electrons (CEs), their therapeutic efficacy should be the highest when deposited in close proximity to the nucleus and DNA. Several studies using the AE-emitters Iodine-123 and Iodine-125 showed indeed an increase in the biological effectiveness at inducing cell death and chromosomal aberrations compared to external x-rays when incorporated into the DNA of the cell (Kassis et al 1990, Slabbert et al 1998, Kassis 2010, Fourie et al 2020). However, more recent studies have suggested that internalisation into the cell nucleus might not be obligatory for AE-emitters and targeting other sensitive cell compartments, such as the cell membrane (Pouget et al 2008, Paillas et al 2016, Bernhardt et al 2021), or cytoplasmic organelles (Santos et al 2024, Spoormans et al 2024), may also induce cell killing. While the AE-emitting radionuclide Terbium-161 has reached the clinical phase after showing promising preclinical results when compared to Lutetium-177 (Borgna et al 2021, Kong et al 2024), novel AE-emitters such as Erbium-165 and Palladium-103 are not well studied. However, the latter radionuclides have potential as more efficient therapeutic agents since they do not have long-range β-emissions and they can be attached to the same biological targeting vectors used to deliver Lu-177, which allow the internalisation of the radionuclide into the cytoplasm (e.g. ligands DOTA-TATE and PSMA-DOTA) or localised on to the cell membrane (e.g. ligands DOTA-LM3 and DOTA-JR11). Further radiobiological preclinical studies are needed to confirm their potential and optimal localisation within the cancer cell (figure 1).
Figure 1. AE emitting radionuclides localised on the cell membrane, internalising into the cytoplasm, and the cell nucleus.
Download figure:
Standard image High-resolution imageMicrodosimetry can give useful insights in this regard, simulating the dose deposition of different AE radionuclides when localised at different sites within the cell, and takes into account the inhomogeneous absorbed dose caused by the variable uptake at subcellular regions and the various emissions of different radionuclides and their unique interaction properties (Dewaraja and Sjogreen-Gleisner 2024, Tamborino et al 2022). The Medical Internal Radiation Dose (MIRD) formalism is widely applied for dosimetry at microscopic and subcellular levels, where the ‘source’ region of activity and ‘target’ dose region can be designated, such as the cell membrane, cytoplasm or nucleus (Stabin and Siegel 2003, Bolch et al 2009). The mean absorbed dose 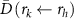 to a target region rk from radioactivity in the source region rh is defined as:
to a target region rk from radioactivity in the source region rh is defined as:

where 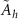 is the integral activity in the source region rh and S is the absorbed dose in the target region rk per disintegration in the source region rh. These so-called S-values in equation (1) may be expressed by
is the integral activity in the source region rh and S is the absorbed dose in the target region rk per disintegration in the source region rh. These so-called S-values in equation (1) may be expressed by

where 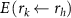 is the total energy deposited in the mass
is the total energy deposited in the mass 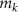 of the target region rk per disintegration in the source region rh, generally expressed in units of Gy/Bq·s; and can be calculated using semi-analytical methods (usually based on the continuous slowing-down approximation) or Monte Carlo methods (modelling of stochastic interaction processes). The goal of this study is therefore to use cellular dosimetry to further investigate and compare novel high-LET AE-emitters, such as Tb-161, Pd-103, and Er-165, amongst others, and look at the impact on the absorbed dose when localised on different cell compartments, particularly the cell membrane, in geometrical models which mimic single tumour cells and micrometastases.
of the target region rk per disintegration in the source region rh, generally expressed in units of Gy/Bq·s; and can be calculated using semi-analytical methods (usually based on the continuous slowing-down approximation) or Monte Carlo methods (modelling of stochastic interaction processes). The goal of this study is therefore to use cellular dosimetry to further investigate and compare novel high-LET AE-emitters, such as Tb-161, Pd-103, and Er-165, amongst others, and look at the impact on the absorbed dose when localised on different cell compartments, particularly the cell membrane, in geometrical models which mimic single tumour cells and micrometastases.
The absorbed energy and dose values were computed using Geant4-DNA (v. 11.1.1) and MIRDcell (v. 3.13). We computed the S-values for various source-target regions—the cell nucleus, cytoplasm, and cell surface (membrane), for uniformly distributed Lu-177, Pd-103 (including the decay of its meta-stable daughter Rh-103 m), Pd-109, Tb-161, and Er-165. We also included well investigated radionuclides such as I-123, I-125, I-131, In-111, and Y-90. The main decay characteristics are summarised in table 1 (supplemental) and figure 2 (Eckerman et al 2008, Duran et al 2020).
Figure 2. Contribution of photons and various electron categories to the energy emitted per decay. Y-90 has been omitted for ease of visualisation.
Download figure:
Standard image High-resolution imageThe Geant4 (Geometry and Tracking version 4) Monte Carlo simulation toolkit is an open-source software developed by CERN for simulating the passage of particles through matter. The Geant4-DNA track-structure expansion aims to provide predictions of biological effects at the cellular level for complex geometrical setups of shielding materials and biological targets (Incerti et al 2009, Bernal et al 2015, Incerti et al 2018).
In Geant4-DNA the event-by-event G4EmDNAPhysicsConstructor_option2 physics list was used with a production cut value of 7.4 eV (the minimum electronic excitation potential of water molecules; below which the residual energy is assumed to be absorbed locally as vibrational energy), ensuring high accuracy in low-energy electron dose calculations. Gamma interactions were also considered and are based on the Geant4 Livermore/EADL97 models, and they are included by default in all G4EmDNAPhysics constructors. The G4RadioactiveDecay model was used with the default photon and electron emission spectra of Geant4. Sufficient decay simulations (⩾500 000) were done such that the standard error of the mean energy and dose were 1% or less.
MIRDcell, a free radiopharmaceutical treatment software, was applied for calculating cellular absorbed doses in single cells and 2D colonies using semi-analytical methods (Vaziri et al 2014, Katugampola et al 2022). The default β-energy spectrum for some common radionuclides used in MIRDcell are obtained from Eckerman and Endo (2008). We investigated the use of both the default emission spectra and the ICRP 107 Nuclear Decay Data, also derived by Eckerman, et al in the dosimetry calculations in MIRDcell (ICRP 2008).
In both Geant4-DNA and MIRDcell, the target cell, nuclei and cell membrane (cell surface) were assumed to be concentric water spheres of unit density. When the source and target region are both part of the same cell (figure 3, left), we termed it the ‘self-dose’; when the particles originate from a source outside of the target cell region, we termed it the ‘cross-dose’ (figure 3, right). In Geant4-DNA we computed the cross-dose contribution to the target from 6 equally spaced and symmetric 1st neighbour cells in the same plane (i.e. a mono-layer); and the final dose contribution was expressed per neighbour cell—the same as in MIRDcell. The radionuclides were randomly distributed in the source region(s) and the dose to the respective target regions were accumulated. In the first model, the cell radius, nuclear radius, and cell membrane thickness were assumed to be 5 μm, 4 μm and 7 nm respectively—similar in shape and dimension to stimulated lymphocytes, Chinese hamster ovarian cells, and rat pancreatic tumour cells in suspension (Humm et al 1994, Fourie et al 2015, Gratte et al 2018). In a second model, the cell radius, nuclear radius, and cell membrane thickness were assumed to be 6 μm, 4 μm and 10 nm respectively to investigate the influence of a larger cytoplasm and cell membrane thickness.
Figure 3. Spherical single cell model (left) surrounded by six 1st neighbour cells in a monolayer (right).
Download figure:
Standard image High-resolution imageThe temporal distribution of the activity is dictated by the cellular pharmacokinetics of the AE-emitting radioligand determined from in vitro experiments, for which data is currently lacking. For simulation purposes it can be simplified to assume that all the radioactivity is located in the aforementioned source regions and remains there undergoing radioactive decay to the stable daughter isotope. When performing dose-response modelling, the cumulative dose to a target region requires the summation of all contributing disintegrations in the possible source regions multiplied by their respective S-value. For example, the dose to the cellular nucleus can be expressed by:
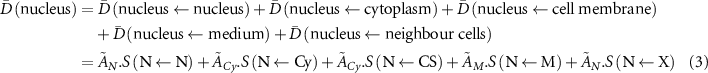
The integral activity in each source region (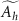 ) is usually determined through in vitro cellular uptake and internalisation experiments and expressed as a fraction of the initially added activity A0. The abbreviations for the nucleus (N), cytoplasm (Cy), cell membrane/surface (CS), medium (M), and cross-dose from neighbour cells (X) are frequently used in literature reporting S-values (target region ← source region).
) is usually determined through in vitro cellular uptake and internalisation experiments and expressed as a fraction of the initially added activity A0. The abbreviations for the nucleus (N), cytoplasm (Cy), cell membrane/surface (CS), medium (M), and cross-dose from neighbour cells (X) are frequently used in literature reporting S-values (target region ← source region).
The self-dose S-values applying the first single cell model (figure 3) were calculated using Geant4-DNA for various target-source regions (supplementary table 2) and for the relative cross-dose contribution per decay per neighbouring cell (supplementary table 4). For example, S(Cy<-CS) = 1.47 mGy Bq·s−1 refers to the mean dose per decay to the cell cytoplasm when I-123 is localised on the cell surface. Some of the S-values are illustrated in figure 4, where the solid bars represent the absorbed doses to the nucleus of the central cell per decay when the selected radionuclides accumulate in the nucleus S(N<-N), in the cytoplasm S(N<-Cy) or on the cell surface S(N<-CS). In addition, the cross-dose from six neighbouring cells were included in figure 4 (outlined white bars).
Figure 4. Self-doses per decay to the nucleus (N) of the central target cell (solid bars) with the additional cross-dose from six neighbour cells (outline) for different cellular localisations of the radionuclides (Cy = cytoplasm, CS = cell surface).
Download figure:
Standard image High-resolution imageIt is observed that Tb-161, Pd-103, and I-125 have the highest S-values for almost every target-source combination. The AE emitter Pd-103 is second to Tb-161, especially when the cross-dose contributions are included, mainly due to their greater CE emissions components compared to I-125. For I-125, which has on average the most AEs per decay, a high local energy deposition is shown in the nucleus when located in the nucleus, but less so for source locations further away (Cy and CS). For both Tb-161 and Pd-103, the dose to the central nucleus is 4–5 times higher when localised in the nucleus, compared to when localised in the cytoplasm or on the cell membrane. Considering the cell surface contribution of 6-neighbour cells, the nuclear dose increased from 1.56 to 3.09 mGy/Bq.s (+98%) for Tb-161 and from 0.98 to 2.24 (+130%) for Pd-103. Upon internalisation in the cytoplasm, the self- and cross-dose S-values showed that the dose to the nucleus was 4-fold higher for Tb-161 and 2-fold higher for Pd-103 when compared to Lu-177.
Figure 5 illustrates the absorbed doses to the different cell compartments when the selected radionuclides are located on the cell surface. Again, I-125, Pd-103 and Tb-161 have the highest S-values for almost every target-source combination. High S(CS<-CS)-values (>30 mGy Bq·s−1) are obtained for most radionuclides, except the β-emitters I-131, Lu-177, and Y-90. For surface-bound radionuclides, the self-dose to the cell membrane substantially exceeds the contribution from neighbouring cell–associated radionuclides, irrespective of the radionuclide used. Interestingly, when located on the cell surface, the membrane dose was similar for Tb-161 and Pd-103; but both 10-fold higher compared to Lu-177. Iodine-125 showed high S-values for nearby targets (i.e. N<-N, Cy<-CS, CS<-CS) due to its large component of AEs but fails to reach targets further away (i.e. N<-Cy and N<-CS) because of its lack of CE.
Figure 5. Absorbed doses per decay to the central target cell (C), cytoplasm (Cy), nucleus (N), and cell surface (CS) (solid) with the additional cross-dose from six neighbour cells (outline) for distributions of radionuclides on the CS.
Download figure:
Standard image High-resolution imagePalladium-109, similar to Lu-177 in its β-electron energy and similar to Tb-161 in its AE and CE energy, also has higher S-values than Lu-177 especially when including the cross-dose contributions from 6 neighbouring cells. Iodine-123, I-131, and In-111 are the radionuclides with the highest photon contribution over the total energy they emit (figure 2) and this is reflected in their low S-values for nearly all target-source combinations. S-values to the nucleus ⩽1 mGy Bq·s−1 were obtained for I-123, I-125, I-131, In-111, Y-90 and Er-165 when located in the cytoplasm or on the cell surface. Interestingly, when Er-165 is located on the cell surface, absorbed doses to the whole cell or cytoplasm are comparable to that of Lu-177, and far exceeds the dose to the cell membrane. Additionally, in all configurations, the dose contribution of neighbouring cell is limited for Er-165 due to its lack of CE and β-particles, being a pure AE-emitter.
To investigate the effect of the cell size and thickness of the membrane, the self-dose S-values applying the larger second cell model were calculated for Lu-177, Pd-103, and Tb-161 (supplementary table 3). The cell radius was increased from 5 to 6 μm and the cell membrane was increased from 7 to 10 nm while including an equal nuclear radius of 4 μm. Our results show that the 3 nm increase in membrane thickness resulted in a ∼40% decrease in S(CS<-CS)-values for Lu-177, Pd-103, and Tb-161.
Finally, the self-dose S-values from Geant4-DNA were compared to those of MIRDcell (supplementary table 2). For example, the I-131 S(Cy<-CS)-value is +8.9% higher than the Geant4-DNA value when using MIRDcell and its default emission spectrum for I-123; and is +4.2% higher when using the ICRP 107 emission spectrum in MIRDcell. By definition, the absorbed dose is dependent on the volume and mass under investigation; therefore, only S-values of similar geometries found in literature are mentioned in tables 2 and 3 for comparison. These include derived values using Geant4 and EGSnrc (Fourie et al 2015, Sefl et al 2015, Seniwal et al 2020; Spoormans et al 2024), from the Geant4 derived GATE application (Tang et al 2021) and PENELOPE (Falzone et al 2015); using MC4 and MCNP6 (Bousis et al 2010, Taborda et al 2016); ETRACK and PARTRAC (Ftáčniková & Bohm, 2000, Li et al 2001); using the manual MIRD formulism (Goddu and Budinger 1997); and using CELLDOSE (Champion et al 2008, Hindié et al 2016, Alcocer-Avila 2021, Hindié et al 2024).
The Geant4-DNA results compare well with the analytical methods of MIRDcell and Goddu, et al (Goddu and Budinger 1997); where the largest differences (up to 30%) were noted when the nucleus is targeted by sources on the cell membrane. Alcocer-Avila, et al found more significant differences (up to 42%), notably for cell surface and intracytoplasmic distributions of Rh-103 m, I-125 and Pt-193 m (Alcocer-Avila 2021). Several studies have reported inconsistencies between the S-values computed using different Monte Carlo codes and those published by MIRD, particularly when the radiation source is distant from the target region, such as is in cytoplasmic or cell surface radionuclide distributions. Hence, it is generally accepted that event-by-event Monte Carlo transport codes provide more accurate cellular S-values than those using semi-analytical methods (Falzone et al 2015, Bousis et al 2010, Alcocer-Avila 2021, Tajik-Mansoury et al 2016). The developers of MIRDcell have also mentioned that the S-values computed with this software can vary significantly from those calculated with Monte Carlo codes for low-energy electron emitters (Vaziri et al 2014). Additionally, we found that the Geant4-DNA values compared better to MIRDcell results using the default emission spectra than when using the ICRP 107 spectrum in MIRDcell, as was employed in Bolcaen et al (2023).
MIRDcell only uses the β-spectrum in their radionuclide radiation data for spheres with radius less than 6.2 mm while contributions from gamma rays, x-rays, neutrons, and recoil energy are not included in the S-value calculations. However, as demonstrated by the IAEA and Howell, et al photon contributions are significant only for self-dose and cross-dose in organ-sized targets, as also noted by (Stabin and Konijnenberg 2000), (Champion et al 2008, Bolcaen et al 2023). This is also reflected in our supplemental Table 4 which shows the contribution of cross-irradiation from neighbouring cells. Our Geant4-DNA values, which include photon emissions and interactions, agree well with those of MIRDcell, which does not include photon emissions and interactions. Therefore, photon emissions and interactions may be omitted for calculations at this microscopic scale.
The Geant4-DNA self-dose S-values compare well within 10% to other Monte Carlo reported doses and most are within 5%, particularly for well-studied source distributions in the cytoplasm, the nucleus, and the entire cell when targeting the nucleus or the entire cell. When compared to other authors, differences ranging from −38.8% to 21.9% are observed for source distributions on the cell surface. A possible reason could be that interactions in the cell membrane are highly dependent on the boundary crossing algorithm and interaction step sizes employed by the simulation package, which could lead to large variations even amongst Monte Carlo codes particularly when having to compromise between accuracy and time of simulations. In Geant4-DNA, the step size for electron interactions are determined dynamically and are inversely proportional to the energy-dependent total interaction cross section. At low energies (<1 keV), where cross sections are large, the mean free path are a few nanometres or less (Incerti et al 2018).
Since in vitro and pre-clinical studies have shown that targeting the cell membrane with AE-emitters may also induce cell killing (Borgna et al 2021, Kong et al 2024), recent efforts shifted to performing microdosimetric evaluations of the cell membrane itself. Because of the very small volume (7–10 nm thickness), this can only be achieved through track-structure Monte Carlo codes; and reference values are not widely available. For all target-source combinations, we found that the S-values of Tb-161 (followed by Pd-103) were greater than that of Lu-177, despite its similar amount of β-electron energy. This aligns with the results of Borgna et al where Tb-161 showed to be 4–5 times more effective at inhibiting tumour cell viability in vitro when labelled to DOTATOC (cytoplasmic localisation) and 102 times more effective when labelled with DOTA-LM3 (membrane localisation) compared to Lu-177. They noted that this increased effectiveness is most likely due to the AE and CE emissions of Tb-161, particularly when localised on the cell surface (Borgna et al 2021, Spoormans et al 2024). This is also reflected in our computed doses: when located in the cytoplasm, the dose to the nucleus was 4-fold higher for Tb-161 and, when on the cell surface, the membrane dose was 10-fold higher compared to that of Lu-177.
The long half-life of Pd-103 (16.99 d) could make it attractive for production and logistics, but the stability of the radiopharmaceutical, low dose rate, and the bio-redistribution of the therapeutic agent might be detrimental to its clinical effectiveness. Although Hindié, et al did not report S-values according to the MIRD formulism, they showed a 10- to 20-fold increase in absorbed dose to the cell membrane compared to the absorbed dose to the cytoplasm for membrane-bound radionuclides Lu-177, Tb-161, and Pd-103 (Hindié et al 2024). We found similar increases, 9-fold for Lu-177, 17-fold for Tb-161, and 30-fold for Pd-103 and Er-165. The increase in dose is largely due to the small volume and mass of the membrane, particularly compared to that of the cytoplasm, however there is a clear dependence and benefit to targeting it using AE-emitters over that of photons or high energy β-emitters like I-131, Lu-177 and Y-90. Hindié, et al also showed that when in the nucleus, Pd-103 can deliver 7- to 10-fold higher nuclear dose and 9- to 25-fold higher membrane absorbed doses than Lu-177 to single tumour cells and cell clusters, where Tb-161 ranked in-between Pd-103 and Lu-177 when comparing absorbed doses. Although when nuclear bound, in our data, Pd-103 ranks between Tb-161 and Lu-177. This could be due to the fact that we did not include the nuclear cross-dose from additional clustered neighbour cells further away from the central cell and its immediate neighbours. However, our single cell self-dose S-values for Pd-103 do agree within 1% to 3% to Hindié, et. al. When nuclear bound our data showed that for Tb-161 the dose to the nucleus was 5-fold higher compared to Lu-177 and for Pd-103 3-fold higher compared to Lu-177. When membrane bound, the membrane dose was 10-fold higher for Tb-161 and Pd-103 compared to Lu-177. Both AE-emitters Pd-109 and Er-165 also showed 5-fold higher membrane dose compared to Lu-177 when membrane bound. However, the short half-lives of Pd-109 and Er-165 (13.7 h and 10.4 h respectively) might only make them useful for research purposes and less practical to implement in the clinic.
As noted above, the cross-dose from a neighbouring cells to the target cell membrane is minimal due to its small volume, whereas cross-dose can be substantial in the nucleus, cytoplasm, and whole cell, particularly from surface-bound radionuclides (figures 4 and 5). The type of particle emissions also affects the cross-dose contribution. Radionuclides such as I-123, I-131, In-111, and Y-90, which predominantly emit photons or high energy β-particles, contribute less to cross dose at the single-cell scale compared to radionuclides like Tb-161, Pd-103, and I-125, which emit CEs along with some higher energy AEs. The contribution increases with the number of cells surrounding the target cell. Alcocer-Avila, et al and Hindié, et al studied the absorbed dose in single cells and clusters, which consisted of six isotropic 1st-neighbour cells and twelve 2nd-neighbour cells (Alcocer-Avila et al 2020, Hindié et al 2024). They found that cross-dose to the central nucleus from the cell surfaces of this cluster made up 67% (Tb-161) and 74% (Pd-103) of the total cell dose, compared to the self-dose alone. The radionuclides contributed more to the cross-dose when located on the cell surface and less when distributed within the nucleus—similar to our results in table 4. As Alcocer–Ávila, et al noted, the superiority of AE emitters like Tb-161 and Pd-103 resides in the very short penetration depths of the low-energy electrons while the cross-dose contributions further than 2-neighbouring cells away would be minimal or even negligible. Enhanced geometrical modelling (e.g. ellipsoid cells) and inclusion of cross-dose effects in clustered cells (1st and 2nd neighbours) are recommended for our follow-up studies for improving dosimetry accuracy, along with the determination of AE and CE emission spectra and their respective dose contributions.
Some additional calculations using Geant4-DNA included absorbed energy per decay in larger water spheres of various sizes (1 μm to 13.4 mm radii) with homogenous radionuclide distributions (supplemental). Our I-125 and I-131 in supplemental figures S1 and S2 match well with similar curves obtained by Li, et al and match their average principle electron energy of 18.3 keV and 183.0 keV respectively (Li et al 2001). As Champion, et al indicated, radionuclides whose absorbed dose reaches a plateau at smaller sphere sizes, such as Pd-103, I-123, and I-125 (∼4.2 mg water sphere, radius ∼1 mm), may be more efficient for treating small tumour targets (including micrometastases, small cell clusters, and isolated tumour cells), compared to higher energy radionuclides, such as Tb-161, Pd-109 and Lu-177, for which a substantial proportion of their disintegration energy escapes and is deposited outside the tumour volume (Champion et al 2008). It is for this reason that both groups highlighted that for clinical dosimetry of small tumour targets (<5 mm in diameter), the knowledge of the percentage uptake per gram is not sufficient and the dose calculation has to be corrected by the absorbed dose fractions to the tumour or its individual cells and their cellular components (Li et al 2001). These can be determined by in depth in vitro studies; where fractionation experiments are performed to define the percentage uptake of a radioligand in different cellular compartments. For example, Borgna, et al compared the uptake of 161Tb- and 177Lu-labeled somatostatin analogues in AR42J cells. The total cell uptake of the radiolabelled somatostatin receptor antagonist was 70%, of which 92% was localised on the cell membrane, 7% in the cytoplasm, and 2% in the nucleus. The use of these uptake and localised fractions are needed in accurate microdosimetry calculations (see equation (1)) and the determination of the relative biological effectiveness of different radionuclides and targeting vectors. For example, using our computed S-values and the DOTA-LM3 data of Borgna et al (2021) the self-dose would be 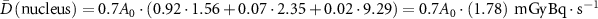 for Tb-161 and
for Tb-161 and 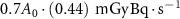 for Lu-177.
for Lu-177.
The Geant4-DNA S-values in this work compared well (within 10%) with other Monte Carlo codes and is believed to provide a more accurate energy and absorbed dose estimations than general-purpose software, particularly regarding interactions and dosimetry in the cell membrane. It was concluded that at a cellular level the dosimetric contribution from emission photons may be omitted.
For the single target cell, the dose to the nucleus was substantially higher with Tb-161 and Pd-103 compared to Lu-177, regardless of the location of the radionuclide in the cell. In the 6 cell micro micrometastasis model, the nuclear cross-dose from the neighbour cell surfaces contributed an additional 98% and 130% for Tb-161 and Pd-103 compared to the self-dose of the central cell. From a dosimetry perspective, the AE and CE emitters could have an improved relative effectiveness when localised on the cell surface, over that of conventional β-emitters. In particular, the membrane dose was 10-fold higher for both Tb-161 and Pd-103 when surface-bound compared to Lu-177.
The authors report no conflict of interest. The authors alone are responsible for the content and writing of the paper.
All data that support the findings of this study are included within the article (and any supplementary information files).
Supplementary data 1 available at https://doi.org/10.1088/1361-6560/ae2b48/data1.
Comments (0)