Access to information on blood flow in various vascular beds is vital across numerous medical fields, guiding clinical decisions and aiding in the diagnosis of cerebrovascular and cardiovascular conditions (Hainsworth et al 2024). However, such data is rarely directly accessible. Available methods are typically indirect and limited in their precision for targeting specific vessels. Quantitative analysis of vascular structure and flow is essential for evaluating patients suspected of cerebrovascular disease (Kalaria 2016). Yet, technical implementation varies based on available tools, time, and expertise. Methods differ in invasiveness, allowing physicians to choose the most suitable option for each clinical scenario (Harik et al 1990). Often, this leads to reliance on highly invasive, such as vascular contrast injection paired with tomographic imaging (Kashani et al 2021).
Transcranial Doppler ultrasonography (TCD) is a noninvasive technique that stands out as a key tool in neurology for assessing vascular reactivity, widely applied in diagnosing cerebrovascular disorders like acute ischemic stroke, vasospasm, assessing brain death or evaluating intracranial pressure in vivo. The latter poses a significant challenge with broad clinical relevance, from traumatic brain injury and subarachnoid hemorrhage to ischemic stroke and brain tumors. The classic approach relies on detecting blood flow asymmetry in intracranial arteries, effective in up to 85% of cases. This method enables rapid, point-of-care flow assessment, which indirectly reflects intracranial pressure. However, in the remaining 15%, an inadequate temporal bone window hampers signal reconstruction and flow estimation from the middle cerebral artery (MCA) (Krejza et al 2007).
The question arises whether non-invasive optical methods, such as Doppler optical coherence tomography (DOCT), used to assess blood flow in the retina, which is easily accessible physically through the transparent ocular tissue, could be used as a proxy of the brain’s vasculature. If positive, an optical evaluation of retinal arterial flow could offer a novel way to assess flow and indirectly gauge intracranial pressure.
Retinal arteries are up to ten times smaller than the MCA (Cavero et al 2017), yet both originate from the internal carotid artery. The ophthalmic artery branches off first, giving rise to the retinal artery within the orbit, while the MCA, a terminal branch, traverses the middle cranial fossa (Paré 1579, Anson 1944). Within the intracranial cavity—housing the brain, cerebrospinal fluid, and cerebral arteries—pressure remains constant but can be altered by breathing exercises (Minhas et al 2018). Breath-holding increases cerebral flow by reducing resistance, whereas hyperventilation decreases flow by constricting intracranial arteries (Cardim et al 2024). These changes also affect the flow spectrum’s shape, reflecting pressure shifts and their sources (Kazimierska et al 2023).
The impact of vascular risk factors, such as hypertension, on cerebral small vessels continues to be explored, with elasticity measures from the internal carotid artery linked to small vessel pathology in the brain’s white matter (Kitagawa et al 2024). Direct measurement of elasticity in retinal small vessels could, therefore, deepen our understanding of the mechanisms underlying cognitive impairment in cerebral small vessel disease (Golzan et al 2014).
The MCA supplies one-third of the intracranial volume, while the eye’s volume is an order of magnitude smaller, with retinal tissue forming only a minor portion (Kauw et al 2019). Despite this difference, retinal arteries share characteristics with cerebral small vessels, making them a valuable model for studying microvascular function. Capturing blood flow in both vascular beds simultaneously offers a unique opportunity to investigate small vessel reactivity. A key metric in this analysis is the pulsatility index (PI) (Michel and Zernikow 1998), which reflects the pulsatile nature of blood flow by comparing its maximum value during systole to its minimum during diastole, quantifying the relative difference between these phases to provide insight into vessel dynamics. Similarly, the resistive index (RI) measures the proportion of flow reduction from systole to diastole (S/D) relative to the systolic peak, indicating downstream vascular resistance. The S/D, in turn, directly compares the peak systolic flow to the diastolic minimum, offering a straightforward gauge of flow variation across the cardiac cycle. These parameters, derived from time series data over multiple cardiac cycles, enhance our understanding of vascular behavior.
A routine TCD assessment of cerebral blood flow is used to evaluate the response to changes in carbon dioxide levels. Increased cerebral flow in response to elevated carbon dioxide levels is well-documented across large studies using techniques like radioisotope angiography (Lassen 1959), bilateral TCD (Mardimae et al 2012), and magnetic resonance imaging (Ziyeh et al 2005). A similar mechanism may exist in ocular arteries, although their smaller volume, diameter, and composition of the eyeball may dampen the local effects of breathing changes. In the MCA, flow volume shifts stem from the reactivity of smaller arteries (below 300 µm) located several branches downstream of the measurement site. In contrast, DOCT directly measures flow in small retinal arteries, where velocity ranges are narrower, driven primarily by changes in diameter rather than resistance.
Studies from the late 1990s suggest that retinal vessels exhibit lower reactivity to carbon dioxide changes compared to cerebral vessels (Harris et al 1995, Schmetterer et al 1997). Doppler laser flowmetry from that period revealed location-dependent responses, showing no significant flow alterations in the optic nerve head (Venkataraman et al 2005, Gallice et al 2017). Later, spatial variability in retinal flow responses to CO2 was observed using optical coherence angiography (OCT-A), providing qualitative insights into flow dynamics by Ashimatey et al (2023).
The DOCT, an advanced form of optical coherence tomography (OCT), can be used for quantitative retinal blood flow measurement, capturing the same physical parameters as TCD does for the MCA. DOCT, an advanced form of OCT, visualizes the retina’s three-dimensional structure, extracting blood cell movement from the OCT signal’s phase. This provides direct access to flow data in specific retinal vessels, yielding parameters like Murray’s metrics (Murray 1926, Riva et al 1985, Pijewska et al 2020) and retinal pulse wave velocity (Spahr et al 2015, Li et al 2018). We detect heartbeat-related changes in velocity profiles by analyzing B-scans over time, enabling the calculation of quantitative indices such as PI, RI, and S/D. Though less studied in small vessels, these parameters are measurable in retinal veins using DOCT (Choi et al 2012, Schmoll and Leitgeb 2013, Huang et al 2016), as well as alternative methods like high-speed CCD photo slit lamps (Koutsiaris 2016) or OCT angiography (Xie et al 2021).
Accurate vessel segmentation is foundational to quantitative DOCT analysis, yet no ‘gold standard’ exists, limiting its clinical use. Techniques for detecting vessel boundaries include histogram filtering (Schmoll and Leitgeb 2013), support-vector machines (Singh et al 2011, Huang et al 2016), and active contour methods (Liu et al 2013). Recently, deep learning approaches have emerged for OCT image segmentation (Lee et al 2017, Shah et al 2018, Wu et al 2019, Tian et al 2021), though none specifically target DOCT vessel segmentation in the human eye.
Although cerebral and retinal vessel activity have been studied separately, concurrent registration has not yet been reported. In this work, we aim to address two technological gaps necessary to address this issue.
The first gap, concurrent recording of vascular responses in both the brain and eye, is addressed by combining intracranial flow measurement using TCD with ocular flow assessment via DOCT.
The second gap, automated vessel detection from DOCT images, is addressed by introducing a novel framework for processing DOCT images, with vessel lumen segmentation as a critical component. Our tailored software framework enables fully automated processing of DOCT data, from acquisition to vessel segmentation and flow quantification, addressing a critical gap in current methodologies and facilitating large-scale vascular analysis. It is designed to advance DOCT as a promising tool for investigating cerebral blood flow through the automated analysis of retinal vessels. Currently, no methods exist for fully automated DOCT data processing, spanning acquisition, vessel detection, and quantification. Our approach overcomes this by automating B-scan preprocessing, alignment, segmentation, and filtration, followed by the detection of retinal microarteries and microveins (diameters of 50–130 µm). It further extracts their area, lumen diameters, and the axial component of blood flow velocity from DOCT images recorded over minutes of heartbeat cycles. With images captured at 95 Hz—surpassing the pulse frequency—we can determine blood flow velocity and lumen values for both systolic and diastolic phases of the cardiac cycle. This enables direct calculation of the PI, RI, and S/D from the recorded time traces.
This integrated approach, not previously reported, when combined with clinically feasible tests such as breath-holding and hyperventilation, allows us to assess how blood vessels in the brain and eye respond concurrently to changes in carbon dioxide levels. Such real-time, synchronized measurements were previously impossible and can provide deeper insights into the regulatory mechanisms of cerebrovascular and ocular blood flow, as well as their interactions.
We recruited ten healthy volunteers and recorded signals during normal breathing, apnea, and hyperventilation. Velocity changes in the MCA are correlated with breath-holding and hyperventilation, while retinal arteries and veins showed no significant statistical correlations. We confirmed that although DOCT can measure blood flow parameters comparable to those obtained with TCD, the flow changes in the eye differ fundamentally from those in cerebral vessels.
This section outlines the key steps of the study, as illustrated in figure 1. Section 2.1 details the experimental setup, while section 2.2 explains the simultaneous DOCT and TCD data acquisition process, alongside respiratory experiments conducted by volunteers to highlight distinct responses in retinal and cerebral vascular beds. Sections 2.3–2.8 present the framework for retinal vessel segmentation from DOCT data, and section 2.9 describes the circulatory parameters derived from DOCT data and methods for comparing them with the TCD signal.
Figure 1. Framework structure and processing pipeline, with references to subsection titles. Refer to the text for details.
Download figure:
Standard image High-resolution image 2.1. Experimental setupDOCT imaging was conducted using a commercial spectral-domain OCT system (RevoNX 130, Optopol Technology Ltd., Poland), while TCD data were acquired with the DopplerBox system (Doppler-BoxX, Compumedics Germany GmbH). The OCT trigger signal was transmitted to the data acquisition card (DAQ NI PCIe-6363, NI National Instruments) to synchronize the probing of the TCD signal. This enabled concurrent acquisition of DOCT images (B-scans) and TCD data at 95 Hz. Details of both imaging protocols are summarized in table 1.
Table 1. Doppler optical coherence tomography (DOCT) and transcranial doppler (TCD) measurement protocols.
DOCT protocol TCD protocol Pixels (z) × A-scans (x) × B-scans (t)1024 × 1024 × 512Power420–450 W m−2Transverse scan range3 mm × 3 mmFrequency2 MHzTransversal/axial imaging resolution12 μm/ 5 μmRange of depth25–52 mmCentral wavelength830 nmGain38B-scan acquisition frequency (B-scan acquisition time)95 Hz (10.5 ms)Sample in time95 Hz (same as DOCT B-scans)A-scan acquisition frequency (A-scan exposure time)100 kHz (9.1 μs) 2.2. Acquisition of Doppler OCT and TCD signals during CO2 breathing testsDOCT images (B-scans) and ultrasonic waves from the TCD probe were simultaneously recorded in the retinal vessels of the human eye and in the MCA. Neither method directly measures total blood flow, capturing only the velocity component parallel to the probing light or ultrasound beam. DOCT B-scans, however, provide 2D profiles of this velocity component for each vessel analyzed. Since both techniques share the same limitation on total blood flow calculation, we rely only on dimensionless parameters for quantitative analysis—specifically, the PI, RI, and S/D—which remain consistent across velocity components.
To compare blood flow velocities in the retinal and cerebral vascular beds, ten volunteers aged 26–45 underwent breathing tests to assess vasomotor responses to artificially induced CO₂ changes in arterial blood under normal air conditions, ensuring simple and natural clinical settings. Data were collected during normal breathing as a baseline. In the apnea test, volunteers exhaled fully and held their breath for 30 s, with OCT imaging conducted for 5 s; this was repeated five times at 10 s intervals. In the hyperventilation test, participants breathed deeply at 1 Hz, guided by a metronome, and after 30 s, DOCT and TCD data were recorded for 5 s. In all cases the spectral shape of TCD velocity was controlled a trained operator to assure the task was done properly by the subject.
The imaging took place at the Oculomedica Clinic in Bydgoszcz, Poland, adhering to the Declaration of Helsinki (World Medical Association 2013). Experimental protocols were approved by the Bioethical Committee of Nicolaus Copernicus University in Toruń, Collegium Medicum in Bydgoszcz (approval ID: KB 87/2021, issued on 16 February, 2021) and all participants provided informed written consent.
2.3. Processing of the OCT dataDOCT is an interferometric technique where the light spectrum, captured by a spectrometer, is modulated by spectral interferometric fringes with a frequency proportional to the optical path difference between the reference mirror and the scattering layer within the sample. Initial processing, including fixed pattern noise removal, wavenumber linearization, dispersion compensation, and spectrum shape correction, yields a complex-valued image line, or A-scan, denoted as I(x,z,t), through Fourier transformation of the corrected spectrum I(x,k,t) (Szkulmowski et al 2016). This process generates a single image line along the z-direction, aligned with the light beam’s propagation. By combining multiple complex A-scans, acquired as a fast optical scanner shifts the imaging beam laterally along the x-direction, a two-dimensional complex-valued DOCT B-scan is constructed:
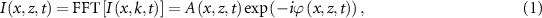
where fast Fourier Transform (FFT) denotes Fourier transformation, z represents the depth coordinate, x indicates the lateral position determined by the x-scanner, A is the OCT signal amplitude, and ϕ is the OCT signal phase. The OCT amplitude encodes the intensity of light scattered from a specific tissue depth, as shown in figure 2(a.2). The signal phase, dependent on the optical path difference between the interferometer arms, enables the measurement of nanoscale displacement by comparing phases from consecutive time points, as illustrated in figure 2(a.1). This principle underpins the phase-based DOCT method, which extracts the axial component 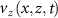 of the blood flow velocity in vessels, according to the following relation:
of the blood flow velocity in vessels, according to the following relation:

Figure 2. Key stages of the data processing pipeline. (a) Aligned OCT images: (a.1) phase B-scan and (a.2) amplitude B-scan, with a green box highlighting a vessel visible in both signal components; subsequent B-scans are cropped to these regions for clarity. (b) Segmentation results: (b.1) single B-scan and (b.2) average of five B-scans, shown after vessel indexing, with a semitransparent mask marking a segmented vessel region. (c) Smoothing of segmented vessel edges: (c.1) lumen radii encoded as intensity values, (c.2) intensity map of vessel lumen radii across all acquired B-scans, and (c.3) the map after median filtering. (d) Final segmentation contours, derived from a parabolic fit to the velocity profile above the noise detection threshold: (d.1) contours overlaid on intensity and velocity B-scans, and (d.2) selected blood flow velocity profile (red) with inner segmentation boundaries marked by blue points.
Download figure:
Standard image High-resolution imageHere, 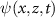 is the unwrapped phase difference between consecutive measurements (Pijewska et al 2019), k is the mean wavelength number, and n is the refractive index of the water. To enhance the signal-to-noise ratio, we employed a variant of DOCT known as spectral and time-domain OCT (STdOCT) (Szkulmowski et al 2008), utilizing more than two complex A-scans to assess phase changes. In our STdOCT implementation, phase shifts were calculated using groups of four A-scans sharing spatial information about dynamic scattering particles in the blood, resulting from dense A-scan sampling relative to transversal resolution (see table 1). These phase changes were unwrapped via a fast phase unwrapping method (Pijewska et al 2019) to allow reconstruction of velocity distribution when phase shifts exceed ±π. After this step, the velocity is integrated over the full vessel cross-section. Further details on DOCT signal processing and phase unwrapping are provided in Pijewska et al (2019, 2020).
is the unwrapped phase difference between consecutive measurements (Pijewska et al 2019), k is the mean wavelength number, and n is the refractive index of the water. To enhance the signal-to-noise ratio, we employed a variant of DOCT known as spectral and time-domain OCT (STdOCT) (Szkulmowski et al 2008), utilizing more than two complex A-scans to assess phase changes. In our STdOCT implementation, phase shifts were calculated using groups of four A-scans sharing spatial information about dynamic scattering particles in the blood, resulting from dense A-scan sampling relative to transversal resolution (see table 1). These phase changes were unwrapped via a fast phase unwrapping method (Pijewska et al 2019) to allow reconstruction of velocity distribution when phase shifts exceed ±π. After this step, the velocity is integrated over the full vessel cross-section. Further details on DOCT signal processing and phase unwrapping are provided in Pijewska et al (2019, 2020).
DOCT processing yields two 3D floating-point arrays containing preprocessed amplitude I(x,z,t) and velocity vz(x,z,t) signal values, hereafter referred to as the velocity data volume and amplitude data volume. Our pipeline’s initial requirement is that the velocity B-scans within the volume are vertically and horizontally aligned, ensuring that corresponding structures in adjacent images occupy the same (x, z) position in the aligned volume. To achieve this, we developed a procedure based on the scale-invariant feature transform (SIFT) algorithm (Lowe 2004), which generates a translation vector for pairs of misaligned images. SIFT features are detected in normalized velocity and amplitude images, and the n best matches between corresponding features in image pairs are identified using the k-nearest neighbors algorithm. The translation vector, derived from the median difference in matched feature locations, aligns each subsequent B-scan with its predecessor. Applied to consecutive B-scan pairs, this process produces groups of aligned B-scans, though gaps may persist where severe artifacts—such as those from involuntary eye movements—disrupt alignment, leaving some groups shifted relative to others. In the second step, we perform group-to-group alignment to correct these inter-group shifts. All B-scans within each group are averaged into a single image, and pairs of consecutive averaged images are processed using the SIFT-based procedure. The resulting translation vectors adjust the groups, ensuring consistent blood vessel positioning across all B-scans. An example of this alignment method applied to consecutive B-scans is demonstrated in supplementary video S1 online.
2.5. Vessel lumen segmentationFollowing the alignment process, the next step is the initial segmentation of the vessel lumen. Segmenting Doppler OCT images with high precision poses significant challenges, particularly in accurately delineating vessel boundaries. Near the vessel walls, where the blood flow velocity is low, contours appear blurred on B-scans, complicating precise boundary identification. Even manual segmentation, where an expert traces vessel outlines on the B-scan, struggles to achieve pixel-level accuracy and proves excessively time-consuming.
To automate lumen segmentation, we employ a U-Net-based neural network model, a convolutional architecture designed by Ronneberger et al for biomedical image segmentation (2015). This encoder–decoder framework incorporates skip connections at each level to enhance detail preservation. Our implementation features pairs of 3 × 3 convolutional layers, each followed by a rectified linear unit activation function and a 2 × 2 max pooling unit on the encoder side, paired with 2 × 2 up-sampling units on the decoder side. The channel progression across consecutive convolutional layers is 64, 128, 256, 512, 1024 in the encoder, reversing to 1024, 512, 256, 128, 64 in the decoder.
The model was developed through a conventional supervised training process. To enhance generalization, we trained it using DOCT data collected specifically for this task from subjects distinct from those in the breathing test experiment. Accordingly, we acquired 16 OCT volume scans, totaling 4064 velocity B-scans (each 1024 × 1021 pixels). For the training and validation datasets, we selected 144 diverse velocity B-scans from the first 12 volumes (from three subjects) and 48 from the subsequent four volumes (from one subject). These B-scans were manually segmented by an expert, and both datasets were expanded in the randomized augmentation process, yielding 15 000 training and 2400 validation samples. Each input velocity B-scan was normalized to a 0–1 range and resized to 256 × 256 pixels via bilinear interpolation, requiring no cropping or manual adjustments. The model outputs a binary segmentation mask, where non-zero pixels denote segmented vessel regions, rescaled to the original size using bilinear interpolation. The training was optimized with the Adam algorithm (Kingma and Ba 2015), employing binary cross-entropy as the loss function and intersection-over-union (IoU) as the performance metric, which evaluates the overlap between true and predicted vessel regions. The initial learning rate was set to 1 × 10−⁵, and a batch size of 1 was used. We empirically set the training duration to 50 epochs, as additional training did not improve IoU values. IoU values, plotted in figure 3, served as an indicator of improving model quality and guided the selection of the best model from those produced during manually supervised experiments.
Figure 3. Intersection over union (IoU) during the training process illustrates the model’s increasing accuracy in comparing true and predicted values across training and validation datasets.
Download figure:
Standard image High-resolution imageThe trained model is integrated into our framework to generate a binary segmentation mask for each velocity B-scan. In the proposed pipeline, we collect B-scans from a single cross-section; thus, to improve segmentation quality, we replace the original velocity image with an averaged image, computed from a velocity B-scan and its ±k neighbors over a time window. We empirically selected k = 5, corresponding to an averaging period of 0.12 s across 11 B-scans. If a B-scan lacks sufficient neighboring images, its segmentation is skipped. Figure 2(b) illustrates the impact of this averaging technique: sample (b.1) shows the segmentation result for a single B-scan, while sample (b.2) displays the enhanced segmentation of the same B-scan averaged over k = 5 neighbors.
2.6. Individual vessel indexingSince multiple vessels may appear within a single data frame, we implement an indexing approach leveraging the initial B-scan alignment. In each binary segmentation image, we identify binary blobs, isolated, homogeneous regions of non-zero pixels, and their central (z, x) coordinates, corresponding to vessels detected by the neural network model. The number of detected vessels typically varies across the volume, influenced by factors such as vessel size, blood flow velocity, or cross-sectional angle, which can render some vessels barely visible. To address this, we determine the maximum number of vessels detected across all B-scans and compute their average (z, x) positions. We then iterate through the segmentation masks, using the aligned B-scans to assign each binary blob an index based on its proximity to the nearest averaged vessel location.
2.7. Angular filtrationThe vessel lumen shape segmented by the U-Net model often appears irregular at its edges, where distinguishing the blood flow velocity signal from noise proves challenging. To address this, we developed a custom filtration method to smooth the initially segmented vessel surface, drawing on three observations about vein and artery cross-sections over time: (1) vessel cross-sections typically exhibit an ellipsoidal shape, (2) their shape evolves smoothly over time, (3) the segmented vessel can be modeled as a 3D object, with 2D cross-sections acquired temporally forming the third dimension.
Figure 2(c) illustrates this process. Each binary segmented vessel edge, figure 2(c.1), is represented in polar coordinates, with the vessel’s center as the origin and its radius sampled at 1° angle increments, plotted as a grayscale image, figure 2(c.2). The array is then smoothed using a 2D median filter (Tukey 1977), employing an empirically selected 7 × 7 square kernel. The filtration result is visualized in figure 2(c.3). This distance map enables retrieval of the smoothed radii to reconstruct filtered binary segmentation images.
As an additional filter, we leverage the expectation of observing a distinct parabolic shape in the blood flow velocity component, rising above the noise level, as shown in figure 2(d). We identify the points where the fitted parabolic profile intersects the mean noise level and adjust segmentation contours accordingly, finding this approach more reliable than segmentation based solely on vessel visibility in B-scans. Vessel boundaries often lack clarity in B-scans alone, whereas the velocity component plot for a single cross-section reveals blood flow down to the phase noise limit, enabling more precise delineation of the vessel lumen. Consequently, this filter expands the contours by up to 10 pixels.
2.8. Vessel features analysisIn the proposed method, for each B-scan, we automatically calculate vessel area (based on segmentation results), diameters (using angular notation from the segmentation), and blood flow velocity at the vessel center (derived from averaged phase images). We compare these values with the reference TCD blood flow velocity signal, as shown in figure 4.
Figure 4. Blood flow-related parameters automatically detected from OCT and TCD. (a) Time series of retinal vessel lumen area, diameter, and velocity extracted from the OCT dataset, automatically segmented using the proposed method, and velocity from the middle cerebral artery recorded via TCD. Colored lines represent raw data with gaps due to eye-motion artifacts or blinks, while gray lines depict the results of FFT-based denoising and gap-filling. (b) Frequency spectrum of OCT and TCD blood flow velocity. (c) Three harmonics of the heartbeat derived from the spectra in (b). (d) Main heartbeat frequency in OCT and TCD velocity signals, scaled to the same amplitude for clarity. (e) Three harmonics of the heartbeat extracted from OCT and TCD, also plotted in gray in the two bottom panels of (a).
Download figure:
Standard image High-resolution imageFigure 4(a) presents the time series of these features for one vessel in an example phase volume. Arrows indicate gaps in the signals due to involuntary eye movements, causing severe artifacts and segmentation failures. These gaps are filled using FFT-based filtering, with the resulting gap-free reconstruction of the feature time series plotted in gray behind each signal.
Figures 4(b)–(e) illustrates the filtration steps. For readability, we analyze only the OCT velocity signal (red) and the TCD velocity signal (black). First, the FFT algorithm is applied to the signals, as depicted in figure 4(b). Next, the heartbeat frequency is identified as the frequency components with the highest amplitudes, shown in figure 4(d), and the phase difference between these components in OCT and TCD is calculated. To reconstruct the gap-free filtered signals, shown in figure 4(e), which enable calculation of systolic and diastolic values for diameters and flow velocities used to determine PI, RI, and S/D, we filter the signal, retaining only the first three harmonics of the heartbeat frequency and the offset.
2.9. Data exploration & visualizationThe procedure outlined in previous sections yields a time series (one value per OCT B-scan) of multiple parameters for each retinal vessel, including maximal, minimal, and mean values of area, diameter, and blood flow velocity, alongside the TCD signal’s blood flow velocity. The OCT-based time series are synchronized point-to-point with TCD-based signals.
All calculated parameters, for both retinal and cerebral vessels, are affected by a systematic error stemming from the unknown orientation of the vessel relative to the measurement plane. This error scales the true parameter values by an unknown factor tied to this orientation. In OCT experiments, the imaging plane, formed by the probing beam and scanning system, is approximately perpendicular to the retinal vessels’ orientation, as most lie parallel to the retina’s surface. However, the precise vessel orientation cannot be derived from the data itself, though it is evident in the ellipsoidal shapes of vessel lumen cross-sections, as seen in figure 5(a). Additionally, Doppler OCT captures only the axial component of the total velocity vector, making it highly dependent on vessel orientation to the imaging plane. Similarly, TCD velocity readings face the same issue, as the artery’s orientation to the ultrasound beam remains unknown. To address this, we adopt a common TCD data processing approach, using unitless parameters—such as the PI, RI, and S/D—to eliminate the scaling factor. These parameters leverage the pulsatile nature of blood flow, where systolic (S) flow represents the maximal value and diastolic (D) flow the minimal value across multiple cycles. We calculate these as follows:



Figure 5. Segmentation validation procedure. (a) Doppler B-scan. (b) Same B-scan as in a. With overlaid manual (violet) and automatic (pink) segmentation borders. (c) Cross-section of the Doppler B-scan along the red line in b. Parabolic distribution of blood flow velocity is visible. Locations of manual and automatic segmentation are shown. (d) The evolution of two selected vessels of different sizes over time is visualized, with the automatically detected border shown as a semitransparent pink layer, and the manual segmentation contour is plotted in violet. Segmentation gaps are marked with red arrows. (e) Rendering of differences between the borders from d.
Download figure:
Standard image High-resolution imagewhere 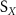 and
and 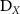 denote the systolic and diastolic values of parameter
denote the systolic and diastolic values of parameter 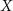 . In subsequent analyses, we focus on OCT velocity (retinal blood velocity,
. In subsequent analyses, we focus on OCT velocity (retinal blood velocity, 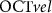 ), OCT lumen area (retinal vessel lumen,
), OCT lumen area (retinal vessel lumen, 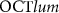 ), and TCD velocity (cerebral blood velocity,
), and TCD velocity (cerebral blood velocity, 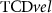 ).
).
The time series are collected and evaluated using an interactive tool we developed to facilitate final data assessment. This tool organizes values into interactive charts, allowing users to select from the listed parameters and signals for comparison, with optional filters for patient ID, breath state during data collection, and phase differences between TCD and OCT velocity signals. Data are displayed across multiple plots simultaneously, with the ability to highlight data point groups in one plot and automatically distinguish them in others. Such comparisons are detailed in the Results section 3.3 , where the tool generated figures 6 and 8.
Figure 6. Aggregated measurements with color-encoded groups, veins coded in blue and arteries in red. Each point represents a single vessel detected and measured in an OCT sample, with point size indicating the delay between two concurrently measured blood flow velocity waves, one from a retinal vessel via OCT and the other from the middle cerebral artery via TCD. (a) Bland–Altman plot of 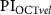 and
and 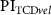 . (b)
. (b) 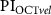 as a function of
as a function of 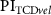 . (c)
. (c) 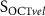 as a function of
as a function of 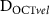 . (d)
. (d) 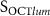 as a function of
as a function of 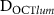 . (e) Bland–Altman plot of
. (e) Bland–Altman plot of 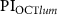 and
and 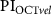 . (f) Bland–Altman plot of
. (f) Bland–Altman plot of 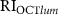 and
and 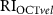 .
.
Download figure:
Standard image High-resolution imageTo support detailed exploration, time series for all acquired signals are accessible for each vessel individually. Additionally, we provide vessel visualizations, utilizing automatically detected vessel locations and segmented lumen contours to create variants of phase and amplitude B-scans, cropped to regions of interest, with and without averaging applied (see supplementary figure S1).
2.10. Statistical analysisIn the statistical analysis, we applied Friedman’s test for all DOCT parameters to account for the following limitations contributing to significant variability in DOCT results: the distribution of retinal vessel sizes is non-normal, with sizes ranging from 50 to 130 μm, multiple vessels originate from a single participant, resulting in dependent data, and, particularly during hyperventilation, increased head movement, coupled with inter-individual breath variations, exacerbates this variability.
For statistical analysis of the MCA TCD data, we employed a repeated measures ANOVA with Greenhouse–Geisser correction, after confirming normality within groups using the Shapiro–Wilk test (positive) and checking variance equality across groups with Mauchly’s test (negative).
Our results are presented across three sections. In section 3.1, we compare our segmentation results with manually segmented data from an expert for validation purposes. Additionally, we detail measurements obtained through our framework: time series of blood flow velocity, segmentation area, and segmentation diameter from OCT, alongside reference blood flow velocity from TCD. We then calculate the PI and RI for the OCT and TCD velocity series, as well as the geometric PI and geometric RI for the OCT segmentation area series. These results are collectively presented in section
Comments (0)